IJCRR - 13(8), April, 2021
Pages: 127-131
Date of Publication: 25-Apr-2021
Print Article
Download XML Download PDF
A Robust Morphological Deep Net Method for Image Segmentation Using Clustering (Retinal Image Segmentation Using Deep Net)
Author: Kaur M, Kamra A
Category: Healthcare
Abstract:Introduction: The segmentation of retinal blood vessel now a day is one of the most important factors which decides the performance of a Computer-aided design (CAD) based system. Segmentation is the process of extracting the region of interest i.e. the disease in the image. The boundaries of retinal blood vessels need to be segmented accurately as an eye surgeon cannot be able to predict the area of disease in case segmentation not done accurately. Objective: This proposed method aims to segment retinal blood vessels using morphological operation which robustly extract the feature. The final image is obtained by using distance-based clustering. Results: The proposed method had shown an accuracy of more than 98.15% and the images are enhanced as the peak signal to noise ratio (PSNR) value is more than 50. Conclusion: The proposed method is efficient in contrast with various existing techniques.
Keywords: Segmentation, Clustering, Morphological, PSNR, MSE, Accuracy
Full Text:
INTRODUCTION
Artificial neural networks with more than two hidden layers are called a deep neural network. Deep neural networks have various architectures depending on their types of connections between layers or operations performed in a layer or unit types in a layer. For example, a multi-layer perceptron has feed-forward connections while a Recurrent Neural Network (RNN) has recurrent connections which provide previous signals to be processed along with the current signal during the training. A Convolution Neural Network (CNN) has convolution layers, performing Convolution between input data and a series of feature detectors. On the other hand, a Deep Belief Net (DBN) has stochastic units and connections between layers are directed from the top layer to the bottom layer.1-3
These networks have been applied to a range of problems from image analysis to language processing. Deep learning has been found very successful at image segmentation, with very extensive examples of object, human or semantic segmentation in natural image.1 To date, there have been limited attempts at organ segmentation in medical images, such as brain part segmentation from MRI images and cell segmentation from microscopy.2,3 Applications of deep networks to retinal vessel segmentation have also started to appear in recent years.3 One key advantage of using a deep network in medical image segmentation can be the adaptation of the method to segment new data, acquired by a different acquisition system, by only retraining the network. In contrast, traditional methods require the adaptation for the segmentation of new data, often entailing the redesign of features according to the new dataset or searching for optimum parameters. On the other hand, the training of a deep network can be challenging in terms of collecting large amounts of labelled data, and this can be viewed as the biggest disadvantage of this method.4-6
MATERIALS AND METHODS
DataSet Used
We have used a clinical dataset obtained from Dr.Ramesh's Super Eye Care & Laser Center, which contains 1800 images, for diabetic Retinopathy.
Algorithm for Proposed Method
(Cross Modality for feature extraction)
Step 1: START
read_image (I)ß Retinal Image ε (D,N,M)
I ß exp (I)
Step 2: Initialize Population à P
For ∀ p generate N feasible solutions for Ii
Step 3: Compute mean (I (Ii) for I ε N
Loop
for i=1 to N àSelection
Select the best two from the population (I1 ´, I2 ´)
Crossover and Mutation ∀ I
Ià[I features ]
Step 4: Set Iß K-classifier
Define Hyperparamter or Set C= empty
Step 5: Divide all pixels with equal distribution
for fold Ki-in K-fold
-
Set Ki as the test set
-
Do distance computation and features selection ∀ I in the loop
-
Loop whilceKfold ≠ NULL
Set Kfoldà K-2 fold
Evaluate Model performance
Compute PSNR ß MSE ε I
End.7,8
In this current work, the cross-modality learning approach is explored for vessel segmentation because it considers the solidarity of label pixels from the same class during the segmentation. Also, this approach suits the nature of the problem as explained below. Fundus image patches can be viewed as noisy versions of vessel masks. Figure 1 shows a fundus image patch and its possible vessel mask.
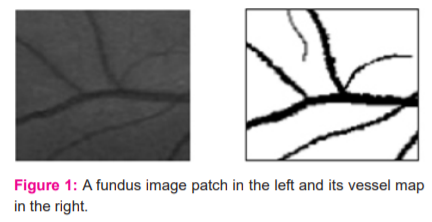

As seen from the figure, the relation between these fundus image patches and their vessel masks is not so complicated, when compared with the relation between the samples of audio and video data in previous applications of cross-modality learning.9,10 In an unrealistic case, even a linear mapping between fundus images and their vessel maps can be possible if the noise level is really low, virtually zero, for fundus images. Because of this similarity, a shared representation learned between fundus images and their vessel masks can react to the characteristics of both data modalities by highlighting the main structures of interest, blood vessels, at the same time.11
The implementation of this approach can be achieved with a generative learning method, such as through using a generative morphological operation (Figure 2). Why a generative learning method is selected can be explained by two reasons. The first is because both generative learning and cross-modality learning require a good representation of the input data. The second is that the features learned during the generative training of a DBN, which can also be called pre-training, can be manipulated to obtain useful features for cross-modality learning.12
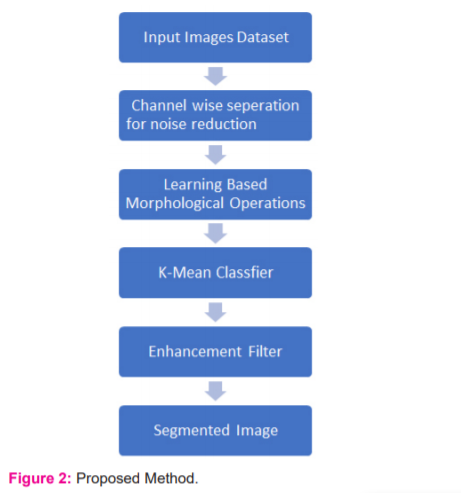
Morphological operations help in smoothening the images and extract the features from the image. The feature extraction is done to preserve the original and the true expected shape of the retinal blood vessel. The various operations are applied as dilation and erosion operations are applied together for edge detection once the threshold is predicted dynamically. The process of dilation is used to separate the pixels from each other. For all the clip window similar pixels the matrix of 1’s is formed and not none matching the matrix of 0’s is formed by which the boundaries can be predicted. The process of erosion will remove the extra boundary pixels to left the user with a crisp idea of the boundary and dimensions. The quality of the mage will be evaluated based on MSE and PSNR. The MSE is the mean square error which is calculated by subtracting the final image from the original. 2,13,14 The PSNR value is computed as:
PSNR = 10log10 [I2 / MSE] …….. (1)
where I range from 0 to 255.
RESULTS AND DISCUSSION
In this section, the performance of the proposed network is evaluated on the CLINICAL dataset. After examining the segmentation performance of the proposed method on the best and the worst-case images, the performance will be compared with that of state of the art methods.
Table I tabulates the overall performance of the proposed network on the CLINICAL dataset concerning the evaluation criteria. Also, the highest and lowest performances based on the maximum and minimum accuracies are shown. As understood from this Table, there is not too much difference between the performance metrics of the best and the worst cases based on accuracy. The proposed network obtained its best and worst performances respectively on the 19th and the 3rd images. These images were also reported as having the best and the worst performances in recent studies using supervised methods.7,15,16 In the Binearization of these vessel probability maps, the average threshold value was found to be 0:1305-0:0432 (average standard deviation). Generated vessel probability maps and binary vessel maps for these images can be found by visual examination of Figure 3.
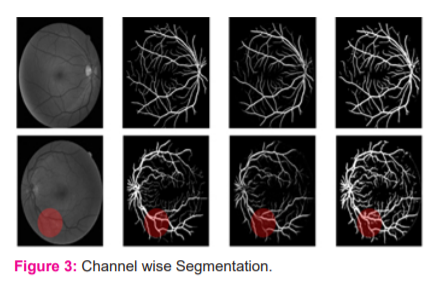
In this Figure, the discrimination of the optic disc from blood vessels in both binary maps (best and worst cases) is performed well, despite the similarity of its border to blood vessels regarding contrast levels. Also, inhomogeneous illumination over fundus images and poor contrast of blood vessels do not seem to cause any disruption in the detection of even tiny blood vessels. On the other hand, the proposed network seems to be sometimes misled by pathologies in fundus images and can sometimes respond to these pathologies as if they are a part of blood vessels. This can be observed in the red circular region in the binary vessel map corresponding to the worst case in the same figure. The proposed network was also seen to mistakenly respond to a fraction of cotton wool spots. Although these responses seem weaker than or almost the same as those of neighbourhood capillaries in the related probability map, some pathological responses appear in the final binary map because of a very low threshold. Also, readers should be aware that the CLINICAL dataset contains random diabetic and non-diabetic fundus images, so it can also be a factor affecting the performance of the network on pathological images. The performance of the proposed network produced a larger AUC value, surpassing the performance of another state of the art deep networks proposed.17-19 Regarding other evaluation metrics, the performance of the proposed method is comparable with the performances of the previous methods. Among them, Author in is the closest to the proposed one: both used a cross-modality learning approach for vasculature segmentation, and both used fully connected networks.20 However, the ways the methods are trained to vary. In the proposed network, the weights in each layer are initialized with weights learned with the probabilistic training of RBMs. However, Li et al. network is a traditional fully connected network with a modification, where the first layer of their network is initialized by the weights learned by training a de-noising autoencoder. The other two layers were initialized by sampling from a normal distribution. Also, the number of hidden layers used in the networks is different (Figure 4).
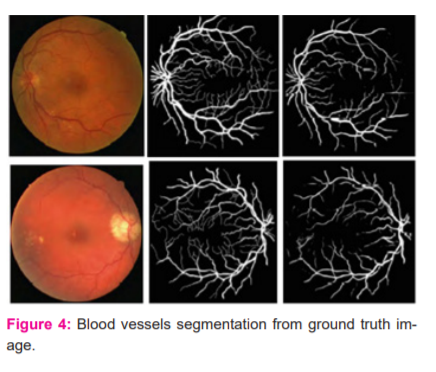
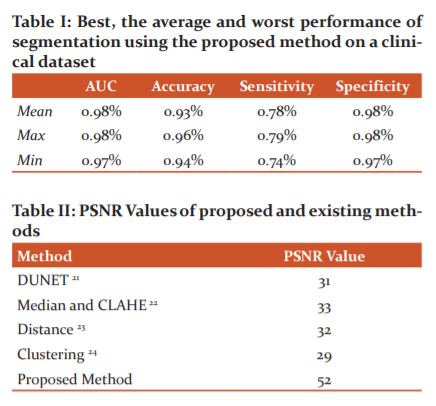
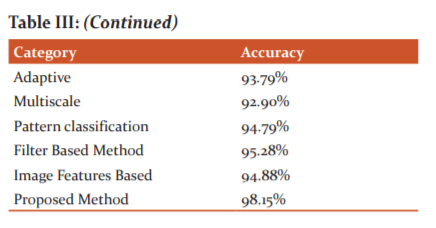
In-vessel segmentation, recently, deep learning approaches have presented more successful results than other approaches.21 This leads me to use a deep network for vasculature segmentation. Although the general tendency in vasculature segmentation is the classification of each pixel in a fundus image as a vessel pixel and a non-vessel pixel, the consideration of spatial connectivity of label pixels in the output of the network has been observed to increase segmentation performance. In this, the spatial connectivity of label pixels is considered by generating vessel masks for given input image patches as a result of complying with the cross-modality learning approach. To improve spatial connectivity of label pixels further, the feature extraction of the proposed is improved with a de-noising, which was found to result in more probability responses to vasculature to the image background.22 Although this effect may not be considered as an improvement regarding segmentation, where probabilities over a threshold would be labelled with the same class, the same effect can be valuable for the robustness of the proposed tracking method (Table I, II and III).
The enhanced segmentation is found to be versatile in terms of applicability to different tasks by slightly modifying its architecture and without significantly changing the way of training the network. When it is used as it is, the network performed comparable or better on the segmentation of low-resolution images, when compared with similar methods.23
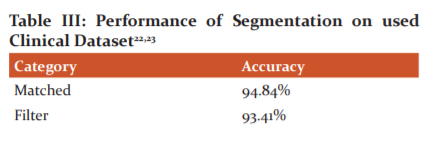

CONCLUSION
As day by day, the number of patients and the necessity of vessel segmentation is increasing. The main reason is to exactly segment the boundaries of the disease to act and treat the patient properly. The segmentation of the blood vessels is done using the RGB separation initially and converting the image in greyscale. The morphological operations are applied to the images to extract the features by which our proposed robust method will segment the retinal blood vessels accurately. The distance-based clustering is used to classify the pixels according to their position, area and intensity to allocate each pixel in the correct cluster. The proposed method had shown an accuracy of more than 98.15% and the images are enhanced as the PSNR value is more than 50. The proposed method is efficient in contrast with various existing techniques. In future, the authors can classify the images based on the segmented retinal blood vessels images.
ACKNOWLEDGEMENT
The authors are grateful to Dr.Ramesh's Super Eye Care & Laser Center, Ludhiana, Punjab, India for providing us with the clinical dataset of retinal images of patients. The authors are also thankful to IKG PTU, Kapurthala, India for providing us with the opportunity to carry out this research work.
This work was supported in my research work
Financial Support: None
Conflict of interest: None
References:
[1] Revathy R. Diabetic Retinopathy Detection using Machine Learning. Int J Eng Res 2019;9(06):8036–8048.
[2] Abdelsalam MM. Effective blood vessels reconstruction methodology for early detection and classification of diabetic retinopathy using OCTA images by the artificial neural network. Informatics Med Unlocked 2020;20:1030-1035.
[3] Khalifa NE, Loey M, Taha MHN, Mohamed HN. Deep transfer learning models for medical diabetic retinopathy detection. Acta Inform Medica 2019; 27(5):327–332.
[4] Takahashi H, Tampo H, Arai Y, Inoue Y, H. Kawashima H. Applying artificial intelligence to disease staging: Deep learning for the improved staging of diabetic retinopathy. PLoS One 2017;12(6):1–11.
[5] Xu K, Feng D, Mi D. Deep convolutional neural network-based early automated detection of diabetic retinopathy using fundus image. Molecules 2019; 22(12):321.
[6] Masood S, Luthra T, Sundriyal H, Ahmed M. Identification of diabetic retinopathy in eye images using transfer learning. Proceeding - IEEE Int Conf Comput Commun Autom 2017:1183–1187.
[7] Pratt H, Coenen F, Broadbent DM, Harding SP, Zheng Y. Convolutional Neural Networks for Diabetic Retinopathy. Procedia Comput. Sci 2016;90(7):200–205.
[8] Carrijo GA, Cardoso F, Ferreira JC, Sousa PM, Patrocínio AC. Image enhancement for blood vessel detection via a neural network using CLAHE and Wiener filter. IEEE Res. Biomed. Eng 2020;36(2):107–119.
[9] Agarwal R, Mahamuni A, Gautam N, Awachar P, Sagar P. Detection of Diabetic Retinopathy using Convolutional Neural Network. Int J Recent Technol Engg 2019;8(4):1957–1960.
[10] Kipli K. Morphological and Otsu’s thresholding-based retinal blood vessel segmentation for detection of retinopathy. Int J Engg Techn 2018; 7(3):16–20.
[11] Lopes UK, Valiati JF. Pre-trained convolutional neural networks as feature extractors for tuberculosis detection. Comput Biol Med 2017;8(9):135–143.
[12] Rahman T, Chowdhury ME. Khandakar A.Transfer Learning with Deep Convolutional Neural Network (CNN) for Pneumonia Detection Using Chest X-ray. MDPI Appl Sci 2020;10:35-41.
[13] Erwin DR. Improving Retinal Image Quality Using the Contrast Stretching, Histogram Equalization, and CLAHE Methods with Median Filters. Int J Image Graph Sig Proc 2020;2:30-41.
[14] Qureshi I, Ma J, Shaheed K. A Hybrid Proposed Fundus Image Enhancement Framework for Diabetic Retinopathy. MDPI Algorithms 2019;14(1)1–17.
[15] Sadikoglu F, Uzelaltinbulat S.Biometric Retina Identification Based on Neural Network. Procedia Comput Sci 2016; 102:26–33.
[16] Alyoubi W L, Shalash W M, Abulkhair M F. Diabetic retinopathy detection through deep learning techniques: A review. Informatics Med Unlocked 2020;20:170-177.
[17] Colomer A, Igual J, Naranjo V. Detection of early signs of diabetic retinopathy based on textural and morphological information in fundus images. Sensors (Basel) 2020;20(4):1–21.
[18] Mateen M, Wen J, Nasrullah N, Sun S, S. Hayat S.Exudate Detection for Diabetic Retinopathy Using Pretrained Convolutional Neural Networks. Complexity 2020;2020(4):1–11.
[19] Islam SM, Hasan MM, Abdullah S. Deep Learning-based Early Detection and Grading of Diabetic Retinopathy Using Retinal Fundus Images. arXiv 2018;23(12):1–13.
[20] Bhupati A. Transfer Learning for Detection of Diabetic Retinopathy Disease Research Project MSc Data Analytics School of Computing National College of Ireland Supervisor?: Dr . Catherine Mulwa.2020.
[21] Jin Q, Meng Z, Pham D, Chen Q, Wei L, Su R. DUNet: A deformable network for retinal vessel segmentation. Knowledge-Based Syst 2019;178(8):149–162.
[22] Raju PV, Varma PK, Nagaraju G. A Dynamic Approach for Exudates Detection in Diabetic Retinopathy Images Using Clustering. Int J Pure Appl Math 2018;119(18):751–765.
[23] Mondal S S, Mandal N, Singh A, Singh K K.Blood vessel detection from Retinal fundus images using GIFKCN classifier. Procedia Comput Sci 2019;167:2060–2069
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License