IJCRR - 5(18), September, 2013
Pages: 01-09
Date of Publication: 25-Sep-2013
Print Article
Download XML Download PDF
BIODEGRADATION OF REACTIVE RED 2 AZO DYE BY BACILLUS LICHENIFORMIS ISOLATED FROM TEXTILE EFFLUENT CONTAMINATED SITE
Author: D. Sudha, R. Balagurunathan
Category: General Sciences
Abstract:A bacterial isolate Bacillus licheniformis was able to degrade reactive red 2 dye, optimally at pH 9, temperature at 37\?C, dye concentration of 50 mg/l at 20% inoculums size. Glucose, NH4NO3 were found to be the best additional carbon and nitrogen sources. The extracellular enzyme from Bacillus licheniformis was studied for dye decolourization potential. Biodegradation was confirmed by analyzing the product using thin layer chromatography (TLC) and gas chromotograpy- mass spectrometry (GC-MS). GC-MS analysis indicated the formation of 2, 4-dichloro-6-[(1H-indazol-5-ylimino)-methyl]-phenol, benzene sulfonamide, 1H indole and urea as final metabolites formed by Bacillus licheniformis. These results indicate the high potential of Bacillus licheniformis to serve as an excellent biomass for the use in reactive red 2 dye removal.
Keywords: Reactive red 2, Bacillus licheniformis, lignin peroxidase, Biodegradation and Gas chromotograpy- Mass spectrometry.
Full Text:
INTRODUCTION
The textile industry plays an important role in the world economy as well as in our daily life, but at the same time, it consumes large quantities of water and generates large amounts of waste water. The chemical reagents used in textile sector are diverse in chemical composition ranging from inorganic to organic molecules (Subhatra et al., 2013). The release of wide range of compounds from industries are creating disturbance to the ecosystem causing climatic changes, reduction of water levels in the ground as well as oceans, melting icecaps, global warming, ozone layer depletion due to photochemical oxidation, etc. (Varsha et al., 2011). Therefore, industrial effluents containing azo dyes must be treated before discharging into the environment to remove the dye toxicity from textile effluent (Rajaguru et al., 2002). Physical and chemical methods (adsorption, chemical transformation, incineration, photo-catalysis, ozonation etc) are not suitable for the removal of recalcitrant dyestuffs, because of high cost, low efficiency and in-applicability to a wide variety of dyes (Dawker et al., 2008).
The microorganisms being highly versatile are expected to develop enzyme systems for the decolourization and mineralization of azo dyes under certain environmental conditions (Pandey et al., 2007). Along with the reductive enzymes, some investigators have demonstrated the oxidative enzymes such as lignin peroxidase, laccase and tyrosinase, in the decolourization and degradation of azo dyes (Bhatia, 2005). Biodegradation using microorganisms are gaining importance as it is cost effective, environmental friendly and produces less sludge (Bella Devassy Tony et al., 2009). In this study, we focused on the isolation and identification of dye-degrading microorganism from textile effluent contaminated site having a decolourizing ability and various intermediates formed have been analyzed during the degradation of reactive red 2 using Gas Chromatography- Mass Spectrometry (GC-MS) techniques.
MATERIALS AND METHODS
Dye
Commercially used textile reactive red 2 was procured from Infra Tex textile industry. The common name of reactive red 2 dye has been used for convenience. Other chemicals used were of analytical grade.
Selection of micro-organism
The effluent sample was collected from Infra Tex textile industry, Perundurai (long 11°13’18.6”N lat 77°39’18.5”E) in Erode district for decolourization studies. The colony was selected on the basis of their ability to form clear zones on this agar plate, containing 50 mg/l of reactive red 2 dye.
16S rRNA sequencing
Amplification of 16S rRNA fragment was performed using16SF (AGAGTTTGATCMTGGCTCAG) and 16SR (TACGGYTACCTTGTTACGACTT). The thermo cycler programme included an initial denaturation of 96°C for 10 seconds and at 94°C for 30 seconds, while it was annealed at 50°C for 5 seconds and extended at 60°C for 4 minutes, these steps cycled a total of 30, while the program was finally extended at 72°C for 10 minutes. The purified products were sequenced by Run 3730, Applied Biosystem 3.0 version (Sudha and Balagurunathan, 2013).
Dye removing efficiency of bacterial culture
For decolourization and degradation study, 250 ml conical flasks containing reactive dyes in mineral salt medium (Moosvi et al., 2007) was inoculated with bacterial isolate. These flasks kept in static incubator at 37°C. After a treatment of 7 days, the decolourized solutions were centrifuged at 10,000 rpm for 4 minutes (Dawker et al., 2008).
Measurement of dye removing capacity
Dye decolourization activity was expressed in terms of decolourization percentage and was determined the absorbance in the supernatent of with drawn samples. Dye removal was calculated according to the equation (Olukanni et al., 2009).
Ao – At
Decolourization (%) = ‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾‾ × 100
Ao
Where:
Ao =Absorbance of the dye solution
At = Absorbance of the treated dyes solution at specific time, t.
Determination of enzyme activities
Bacillus species was grown in 100 ml nutrient broth at 30°C for 24 hours and centrifuged at 6000 rpm for 20 minutes. These cells were suspended in 50 mmol-1 potassium phosphate buffer (pH 7.4), maintaining temperature at 4°C. This extract was used as an enzyme source without centrifugation (Dawker et al., 2008).
Oxidative enzyme during decolourization
Lignin Peroxidase Activity Assay
Lignin peroxidase activity has been performed at 40°C using CaCO3 as a substrate. The assay mixture contains 0.1M citrate/phosphate buffer pH 4.0. The oxidation of CaCO3 led to an absorbance increase at 420 nano meter. One unit of the enzyme activity was defined as the amount of enzyme required to produce 1molar of oxidized product/ minutes under the assay conditions (Dawker et al., 2008). Lignin peroxidase was determined by monitoring the formation of propanaldehyde. The two chemical reagents were used for propanaldehyde estimation such as 2, 4-dinitrophenylhydrazine reagent and Schiff reagent (Bansal Raj, 2009). Cell supernatant and Schiff reagent was added and the colour change was noted after two minutes recommended by schiff test.
2, 4 - Dinitrophenyl Hydrazine Test
Cell supernatant was collected into reaction tube and 20 drops of 2, 4-dinitrophenyl hydrazine was added. Then precipitate formation was noted, if precipitation was not occurred, the next step was carried out. The reaction mixture was rinsed, it indicates the acid removal and recrystallized the product using ethanol (5 ml) and allowed to dry. Then to heat for few minutes and again 2, 4-dinitrophenyl hydrazine was added. Finally colour change was noted.
Biodegradation assay through TLC
After complete decolourization by bacterial isolate at seven days of incubation period, the decolourization medium was centrifuged at 10,000 rpm for 4 minutes and supernatant extracted with ethyl acetate (Dawker et al., 2008). The extract and the aqueous phases were separately evaporated in a rotary evaporator. The concentrated extracts were dissolved in methanol and used for thin layer chromatography (TLC). The mobile phase for the organic extracts was methanol: Hexane (4:1) the bands of aromatic compounds were observed under UV light and spotted then it was scrapped and mixed with methanol; further gas chromatography- mass spectrometry was carried out.
GC –MS analysis
Gas chromatography is one of the most versatile and ubiquitous analytical technique in the laboratory, it is widely used for the identification of organic compounds, when coupled with mass spectrometry as a detection system (Christian, 2009). GC-MS analysis was performed using a Thermo DSQ II Mass spectrometer fitted with GC Trace Ultra Version 5.0 at temperature programming mode with DB 35-MS capillary non polar column. The initial column temperature was held at 50°C then increased linearly to 260°C at 10°C/ minutes. Helium was used as a carrier gas with a flow rate of 1.0 ml/ minutes. Identification of degradation products were made by comparison of retention time and fragmentation pattern with known reference compounds as well as mass spectra in the NIST spectral library.
RESULTS
Isolation, identification of dye decolourizing bacterium
The bacterial strain was selected based on their ability to form a clear zone on nutrient agar plates containing reactive red 2. Among the seven isolated bacterial strains, one strain was selected based on its ability to form high dye decolourization zone on agar plate. The morphological and molecular identifications were performed. This selected strain was identified to be Bacillus licheniformis by 16S rRNA sequencing. The sequence of the 16S rRNA gene of Bacillus licheniformis is available with accession number KC-866382 (Sudha and Balagurunathan, 2013).
Decolourization under different culture conditions
Decolourization of reactive red 2 was studied at different culture conditions. Maximum decolourization was observed at glucose concentration of 10 g/l there was 82%, ammonium nitrate at 3 g/l there was 67% decolourization and dye concentration of 50 mg/l, 85% was achieved at 20% of inoculums. In addition to this, at pH 9 there was 80% colour loss, optimum temperature 37°C there was 79% decolourization after seven days of incubation at static conditions. This indicates that the presence of easily catabolisable C and N sources in the medium enhanced the efficiency of dye degradation. pH 9 and temperature 37°C was found to be most suitable for maximum decolourization.
Enzyme Activity
Estimation of lignin peroxidase enzyme
In order to gain additional insight into the decolourization mechanisms, screening of oxidative enzyme activities were also monitored. The activity of extracellular lip was quantified as 19.8 units per gram substrate in Bacillus licheniformis under the optimized growth condition at 7th day of incubation and qualitatively performed by Schiff Test and 2, 4-Dinitrophenylhydrazine Test.
Schiff Test
Meganta colour was formed in the reaction; it indicates presence of Propanaldehyde (figure 1).
2, 4-Dinitrophenylhydrazine Test
Brown colour precipitate was formed. It indicates presence of Propanaldehyde. So these two methods denote the presence of lignin peroxidase enzyme (figure 2).
Thin layer chromatography
The comparison of TLC chromatogram of the media extracted by the organic phase after decolourization by the Bacillus licheniformis under UV light showed that the decolourized sample had three addition bands, which might originate from the dye metabolites. Aromatic amines are the usual decolourization products of azo dyes that appear in the organic phase extract.
GC-MS analysis
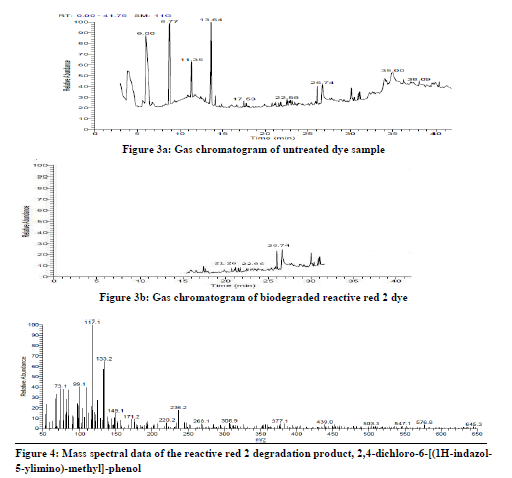
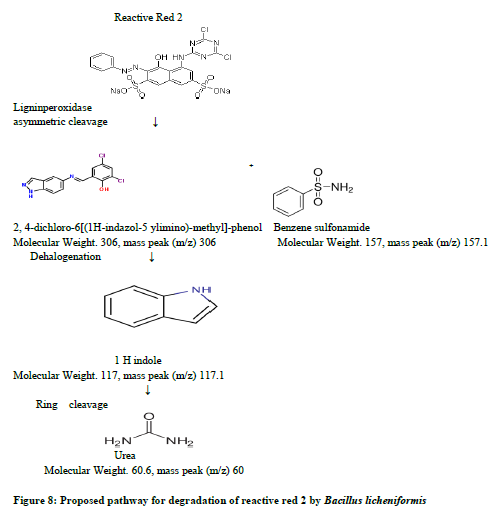
In order to verify degradation products formed during dye decolourization by Bacillus licheniformis, the gas chromotograpy- mass spectrometry analysis was carried out. The untreated textile dye reactive red 2 showed nine peaks in its chromatogram. The compounds analyzed for these peaks were found to be toxic products present in the untreated raw dye sample (figure 3a). Biodegradation analysis showed a major reduction in the entire organic compound and the peaks that were observed was reduced to three to four to significant extent (figure 3b). The pathway is proposed in degradation of reactive red 2, figure 8 showing various steps involved in dye degradation.
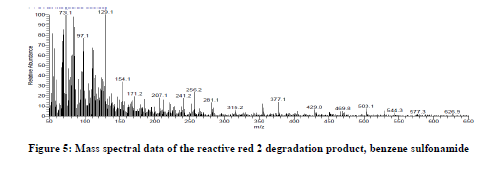
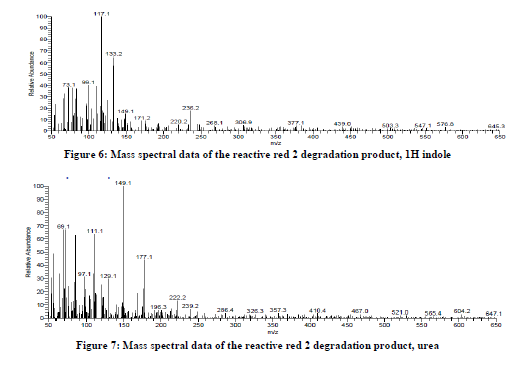
According to our proposed pathway, the peroxidase catalyzed initially the asymmetric cleavage, resulted in the intermediate product which was identified as 2, 4-dichloro-6-[(1H-indazol-5-ylimino)-methyl]-phenol with retention time 26.74 minutes and a mass peak of 306 (figure 4), benzene sulfonamide with retention time 22.96 minutes and a mass peak 157.1, (figure 5) supports the oxidative asymmetric cleavage of reactive red 2. The reduction of 2,4-dichloro-6-[(1h-indazol-5-ylimino)-methyl]-phenol giving rise to the intermediate product in this reaction as 1H indole at retention time 26.74 minutes and a mass peak 117 (figure 6). Further ring cleavage reaction might be followed giving rise the intermediate urea at retention time 21.26 minutes and a mass peak 60.6 (figure 7) in this pathway. Thus salient feature of this mechanistic proposed pathway are the release of azo linkage and formation of phenyl radicals, urea as intermediate which indicates that Bacillus licheniformis have potential to degrade the dye reactive red 2.
DISCUSSION
In this study we examined the good decolourization ability exhibited by Bacillus lichenifornmis isolated from textile effluent. Bacillus licheniformis showed decolourization of reactive red 2 at static condition. Optimization of parameters such as carbon source, nitrogen source, carbon + nitrogen source, pH, temperature, inoculums size, dye concentration in reactive azo dye was carried out in this study. The pH plays great influence in decolourization of reactive red 2 dye. The optimum pH was found to be 9 for maximum removal of dye. These present findings are closely similar with El-Sersy (2007). The range of activity on decolourization of reactive red azo dye with 37°C was high with Bacillus licheniformis. present findings were similar to that Ponraj et al., (2011) they reported that the range of activity on decolourization of orange 3 R with 37°C was 78.57%, Bacillus sp. was found to be the most effective decolourizer. Dyes being deficient in carbon sources the biodegradation of dyes without any extra carbon source is very difficult (Senan and Abraham, 2004). Therefore, in this study, glucose was used as carbon source supplemented in the mineral salt medium.
In this present study Oxidative biodegradation takes place upon action of enzyme lignin peroxidase. In previous studies Moharrery et al., (2012) reported that Halomonas strain D2 was able to degrade Toluidine Red due to its enzymatic activity. In this study the addition of lignin peroxidase inducer CaCO3 to the culture medium of microorganism may enhance lip production and facilitate its utilization, in previous study Dawker et al., (2007) reported that the addition of lip and laccase inducers such as CaCO3, indole, veratrole, vanilline and toludine to the culture medium of microorganism may enhance lip and laccase production. Despite the fact that untreated dyeing effluent may cause serious environmental and health hazards, thus it is of concern to analyse the biodegradative product of the untreated and treated effluent sample. According to our proposed pathway, Low molecular weight aromatic compound such as phenol, 1H indole and urea were produced from the degradation of reactive Red 2 by Bacillus licheniformis, our results similar to that Soundararajan et al., (2012). In this study ring cleavage was identified as the final product, this present findings similar to that Jun lin et al., (2010). Hence, this study indicates that the Bacillus licheniformis have a potential to degrade reactive red 2 dye.
CONCLUSION
The results of these findings suggest a great potential for bacteria to be used in decolourization of real dye waste waters. Interestingly, the bacterial species used in carrying out the decolourization in this study are isolated from the dye industry effluents. Thus biological processes that are simple, fast and economical can be adopted by textile and dyeing industries as an effective alternative for treating their wastewater.
ACKNOWLEDGEMENT
The authors express genuine thanks to the Vice-Chancellor and the Registrar, Periyar University, Salem, for providing research facilities. The authors are also grateful to authors / editors / publishers of all those articles, journals and books from where the literature for this article has been reviewed and discussed.

References:
- Bella Devassy Tony, Dinesh Goyal, Sunil Khanna. Decolourization of textile azo dyes by aerobic bacterial consortium. International Biodeterioration and Biodegradation 2009; 63: 462-469.
- Bhatia S C. Text book of biotechnology. Atlantic Publishers and Distributors 2009; 394-397.
- Dawker V V, Jadhav U U and Govindwar. Biodegradation of disperse textile dye brown 3REL by newly isolated Bacillus sp.vus. Journal of Applied Microbiology 2008; 105: 14-24.
- El-Sersy N A. Bioremediation of methylene blue by Bacillus thuringiensis 4G 1: application of statistical designs and surface plots for optimization. Biotechnology 2007; 6(1): 34-39.
- Gary D, Christian. Analytical chemistry. Pashupati Printers Pvt Ltd 2009; 574.
- Jun lin, Xingwang zhang, Zhongjian li, Lecheng lei. Biodegradation of reactive blue 13 in a two-stage anaerobic / aerobic fluidized beds systems with a Pseudomonas sp isolate. Bioresource technology 2010; 101: 34-40.
- Moharrery M, Otadi A, Safekordi R, Amiri and Ardjmand M. Biodegradation of Toluidine Red, an oil Soluble Azo Dye, With Halomonas Strain D2. World Applied Sciences Journal 2012; 18 (8): 1065-1072.
- Moosvi S, Kher X and Madamwar D. Isolation, Characterization and decolourization of textile dyes by a mixed bacterial consortium JW-2. Dyes and pigments 2007; 74: 723-729.
- Olukanni O D, Osuntoki and Gbenle G O. Decolourization of azo dyes by a strain of micrococcus isolated from a refuse dump soil. Biotechnology 2009; 8(4): 442-448.
- Pandey A, Singh P and Iyengar L. Bacterial decolourization and degradation of azo dyes: a review. International Biodeterioration and Biodegradation 2007; 59: 73-84.
- Ponraj M, Gokila K and Vasudeo Zambare. Bacterial Decolourization of Textile Dye – Orange 3R. International Journal of Advanced Biotechnology and Research 2011; 2(1):168-177.
- Raj K, Bansal. Laboratory manual of organic chemistry, New Age International Pvt Ltd Publishers 2009; 80.
- Rajaguru P, Vidya L, Baskarasethupathi B, Kumar P A, Palanivel M and Kalaiselvi K. Genotoxicity evaluation of polluted ground water and in human peripheral blood lymphocytes using the comet assay. Mutation Research 2002; 517:29-37.
- Senan R C and Abraham E T. Bioremediation of textile azo dyes by aerobic bacterial consortium. Biodegradation 2004; 15: 275-280.
- Soundararajan N, Gopi V, Akilesh upgade and Nazma Begam. Annals of Biological Research 2012; 3(3): 1773-1778.
- Subhathra M, Prabakaran V, Kuberan T and Balamurugan I. Biodegradation of azo dye from textile effluent by Lysinibacillus sphaericus. Sky Journal of Soil Science and Environmental Management 2013; 2(1): 1-11.
- Sudha D and Balagurunathan R. Effect of process parameters on anaerobic decolourization of reactive azo dyes using Bacillus licheniformis isolated from textile effluent contaminated site in Perundurai, India. International Research Journal of Environment Sciences 2013; 2(8): 37-43.
- Varsha Y M, Naga Deepthi C H and Sameera Chenna. An Emphasis on Xenobiotic Degradation In environmental clean up. Journal of Bioremediation and Biodegradation 2011; S (11).
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License