IJCRR - 3(11), November, 2011
Pages: 41-48
Print Article
Download XML Download PDF
NON-DESTRUCTIVE GENETIC SAMPLING IN TWO CYPRINID FISH SPECIES OF SOUTHERN PART OF
WESTERN GHATS-INDIA
Author: Raja. M, Nandagopal. S , Malaiammal. P
Category: General Sciences
Abstract:Non- destructive genetic sampling for DNA isolation in two cyprinid endemic fishes of the
species Danio aequipinnatus and Puntius tambraparniei was attempted. The total genomic
DNA was isolated from fin clips, scales, liver and muscles by using Phenol: choloroform
method by different storage method like Ethanol/EDTA and air dry in the different cell lysis
buffer like TNES- urea and TNES. In Puntius tambraparniei showed higher quantity of
genomic DNA (402.36\?g) in the liver tissues by using the TNES cell lysis buffer in the ethanol/
EDTA storage method. Danio aequipinnatus also showed the high quantity of genomic DNA
(415.26\?g) in the muscle tissue by using TNES cell lysis buffer in the Ethanol/ EDTA storage
method.
Keywords: Non-destructive, genetic sampling, DNA extraction, fin clips, scale, muscle, cyprinid fish.
Full Text:
INTRODUCTION
In recent years many vertebrate species are at risk and their methods of conservation is possible without destruction and based on this DNA based studies gain importance to gain information on the diversity and population analysis (O‘Brien 1994). DNA analysis will help to study the phylogeny, determination of population size and level of genetic polymorphism within and between populations. Liver and muscle tissues are used for DNA analysis and this method implies the sacrifice of the animals and hence it called as destructive method. The non-destructive sources of DNA are hair, faeces, urine, shed feathers, snake skin, sloughed whale skin, eggshells and even skulls. However, this method usually results in a low quantity and poor quality of DNA. Non-destructive sampling also includes the use of blood, skin and scales (Hilsdorf et al. 1999). Muscle tissues and blood samples are used for DNA isolation without sacrifying the fish especially large population of threatened fishes. (Cummings et al. 1994) and (Estoup et al. 1996) Even this can be overcome by using fins and scales. Hence an attempt has been made to compare the destructive method of extracting DNA from muscle and liver tissues and a non-destructive method of extracting DNA from fins and scales of two freshwater fish species of India. One is an ornamental fish Danio aequipinnatus which has the natural distribution in clear water streams/rivers of India, and another species is Puntius tambraparniei an endemic species confined to its distribution range to Tamiraparani river basin alone. (Silas 1953, Arunachalam and Sankaranarayanan 2000).
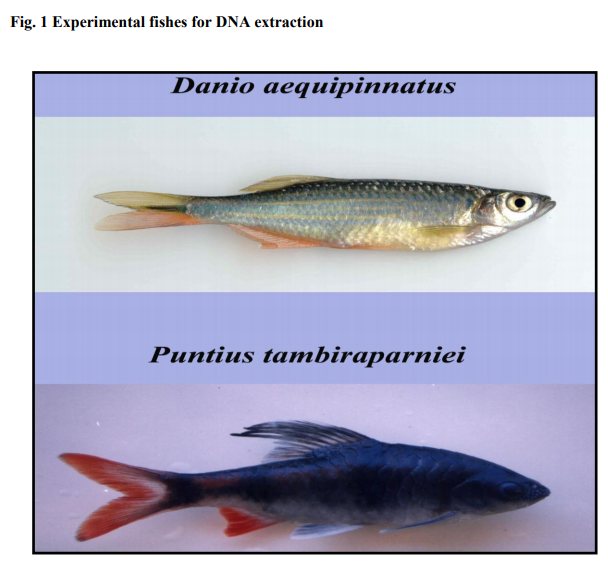
MATERIALS AND METHODS Sample Collection: Cyprinids species Danio aequipinnatus, Puntius tambraparniei were collected from river and cannels, middle reach of the rivers Gadana and Ramanadhi which are the two such basins of the Tamiraparani river system. Fish sampling was performed by using cast net and gill nets.
Extraction of DNA
The genomic DNA was isolated by phenolchloroform method based on Sam brook et al. (1989). DNA was achieved from fish caudal (or) anal fins and from fish scales by changing the protocol previously described for tissue preservation and DNA extraction from muscle (or) liver (Asahida et al. 1996, Sam brook and Russell 2001). Total DNA was obtained from individually belonging to two different fish species Puntius tambraparniei and Danio aequipinnatus.
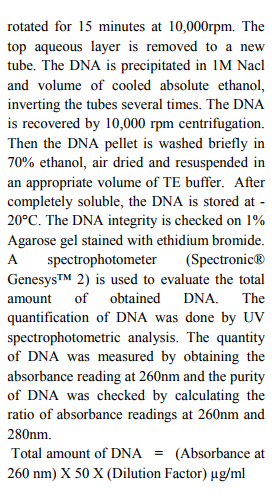
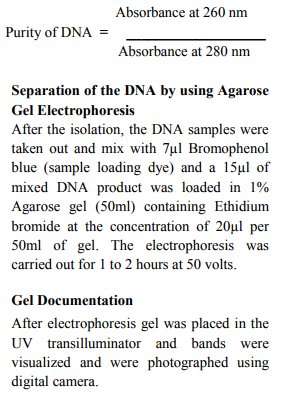
Approximately 100-500 mg of fins (1-2cm2 ) or scales are initially stored in 95% ethanol- 100µl EDTA pH 8.0 (Dessauer et al. 1996). 100 mg freshly airdried samples Fins and scales are collected from Puntius tambraparniei and Danio aequipinnatus. The samples are cut into small pieces using sterile blade. The samples are placed in 4ml of a TNES, TNES- urea -digestion buffer inside of 15 ml tube. 30µl of RNase (10mg/ml) is added to the tube. The samples are incubated at 42°C for 1h. After this period, 30µl of Proteinase K (10mg/ml) is added and tissues are maintained at 42°C for at least 10 hours. The DNA is then isolated by adding 4 ml of phenol: choloroform: isoamylalcohol (25:24:1) to the tubes. After inverting the tubes


RESULTS
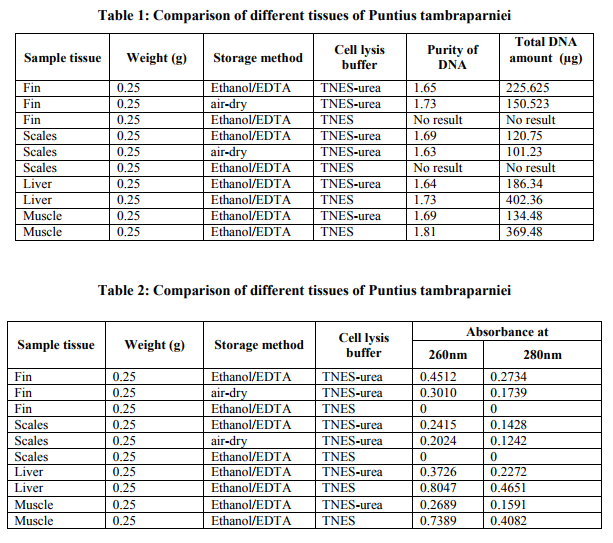
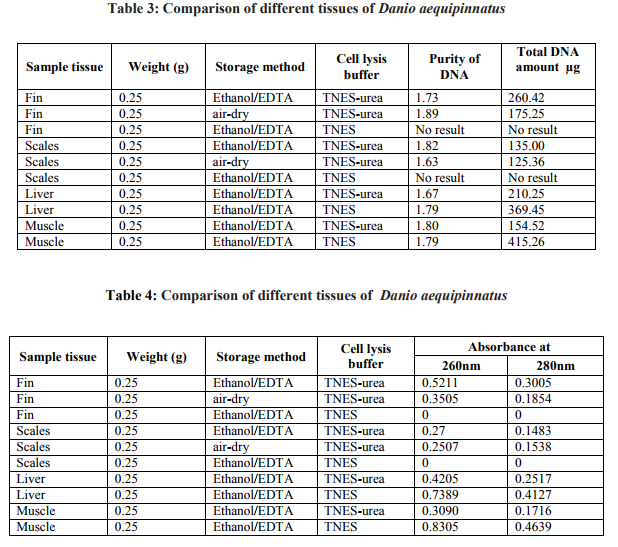
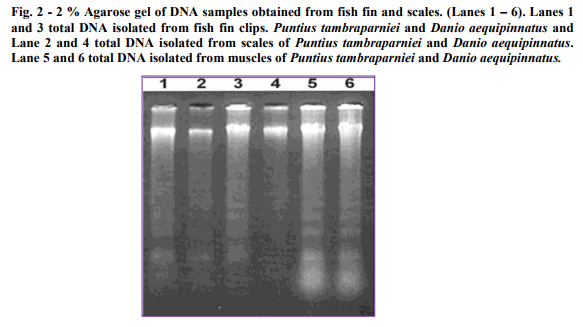
Fin samples (n=3) of Puntius tambraparniei showed higher quantity of genomic DNA using TNES/urea in Ethanol/EDTA and TNES/urea in air dried methods, 225.625µg/ml and 150.23µg/ml respectively; where as in TNES without urea solution shows the absence of DNA. Scale samples (n=3) of Puntius tambraparniei showed higher quantity of genomic DNA using TNES/urea in Ethanol/EDTA and airdried methods, 120.75µg/ml and 101.25µg/ml respectively; where as in TNES without urea solution shows the absence of DNA. Liver samples (n=2) of Puntius tambraparniei showed higher quantity of genomic DNA using TNES method than TNES/urea, 402.36µg/ml and 186.34µg/ml respectively. Muscle samples (n=2) of Puntius tambraparniei showed higher quantity of genomic DNA using TNES method than TNES/urea, 369.48µg/ml and 134.48µg/ml respectively (Table. 1-2). Fin samples (n=3) of Danio aequipinnatus showed higher quantity of genomic DNA using TNES/urea in Ethanol/EDTA and TNES/urea in air-dried methods, 260.42µg/ml and 175.25µg/ml respectively; where as in TNES without urea solution shows the absence of DNA. Scale samples (n=3) of Danio aequipinnatus showed higher quantity of genomic DNA using TNES/urea in Ethanol/EDTA and TNES/urea in air-dried methods, 135.00µg/ml and 125.36µg/ml respectively, where as in TNES without urea solution shows the absence of DNA. Liver samples (n=2) of Danio aequipinnatus showed higher quantity of genomic DNA using TNES and TNES/urea method, 369.45µg/ml and 210.25µg/ml respectively. Muscle samples (n=2) of Danio aequipinnatus showed higher quantity of genomic DNA using TNES and TNES/urea method, 415.26µg/ml and 154.52 µg/ml respectively (Table. 3-4). Most of the isolated DNA of the two species showed no sign of degradation and the spectrophotometer comparison of absorbance at 260 - 280nm provided a DNA/ RNA relationship of (1.6-1.9) indicating good DNA quality. The DNA concentration ranged from 25-500ng/µl, with on average concentration 200ng/µl and the obtained DNA volume (approximately 1ml) was high enough to be employed on several molecular experiments. Although the present methodology was applied on samples of two fish species, similarity in fin (or scales) anatomy suggests that the technique will work on samples of different taxa.
DISCUSSION
As stated by some authors (Chen et al. 1995; Strassmann et al. 1996; Pinto et al. 2000), tissue homogenization in liquid nitrogen can be an efficient method to isolate significant amounts of DNA, especially on hard consistent tissues. However, in our experiments the use of nitrogen maceration with fins and scales did not give any further improvement in the DNA isolation. Better results were achieved in the present study by mixing the scales or small pieces of the fins with a cell lysis solution containing urea. The initial 8M urea concentration of the buffer, suggested by Asahida et al. (1996), was gradually decreased to 4M, which allowed a better preservation of the material and a non-degraded isolated DNA. Urea treatment seems to be a necessary step to breakdown hard tissues such fins and scales, since it is quite denaturing for protein and at least it disrupts most likely any protein multicomplexes. Another improvement on the DNA isolation process was provided by a pretreatment with RNase, which allowed us to obtain DNA samples with lower quantities of RNA that could interfere in the accurate DNA quantification and on further amplification procedures. The concentration and time/temperature for Proteinase K incubation were also very important to obtain high-quality DNA. The use of lower concentrations of this enzyme resulted in poor quality-DNA, as it failed to completely digest the tissue. A better dissociation of the tissues was also obtained when the digestion was done at 42°C. Higher incubation temperatures (50°C or more) were inefficient and temperatures lower than 42°C resulted in a partially digested tissue. Experiments using a final concentration of 0.075 mg/ml of Proteinase K provided tissues that were totally digested after a 10 hours-incubation. Less-time incubation was not efficient. After tissue digestion, a phenol:chloroform:isoamyl alcohol purification step was utilized, as suggested by Taggart et al. (1992) and Sambrook and Russell (2001). The use of phenolchloroform proved to be essential to obtaining pure DNA samples from fish fins and scales. Crude extractions could result in a DNA contaminated with proteins that may not be stable for long-term storage. However, repeated DNA extractions with phenol-chloroform were not necessary. Single and double washes gave same results, avoiding protein residues. The described technique was applied on air-dried and ethanol/ EDTA-fixed fin clips and scales and also on ethanol/ EDTApreserved liver and muscle tissues. In addition, DNA samples were also obtained from liver and muscle using nitrogen maceration and by the use of a digestion buffer without urea, as described in Sambrook and Russell (2001). The use of a lysis solution without urea showed to be not appropriate for fin clips and scales. It is evident that fins and scales, represent a DNA source as suitable as other tissues and also the DNA amount isolated from the fins was also high, when compared to the amount obtained from liver or muscle. Therefore, the extraction of DNA from fish fins or scales offers an extremely a positive alternative to conventional DNA isolation techniques, representing a minimally destructive sampling approach. An adequate preservation of tissue samples is a prerequisite in field locations and for long-period analyses. Despite the successful isolation of DNA from different tissues of Puntius tambraparniei and Danio aequipinnatus, some differences were observed in relation to the material storage. Samples of fins and scales preserved on ethanol/EDTA proved to be more suitable as a DNA source, when compared to airdried samples stored for 1 to several days. Preservation of nucleic acids depends primarily on the inhibition of tissue nucleases and denaturation, which can be achieved with EDTA and ethanol, respectively (Dessauer et al. 1996). The ethanol/EDTA storage solution also permits to maintain the softness of the tissue, facilitating its further dissociation in the digestion buffer. However, the long term storage of tissues on TNES- Urea buffer, as suggested by Asahida et al. (1996), was not appropriate for fins or scales due to a high DNA breakdown. The protocol outlined in this work offers a cost-efficient and suitable alternative to conventional DNA isolation techniques, representing a non-destructive sampling approach to isolate high-quality DNA from fish. The total amount of isolated DNA (25–500µg) is also sufficient for several other molecular procedures that often demand more DNA than the pictogram range. It can be considered that a genetic stock of several fish species could be easily achieved by using the described methodology.

References:
1. Arunachalam, M. and A. Sankaranarayanan (2000). Puntius arulius tambraparniei - An ornamental stream fish endemic to Tamiraparani River basin, South Tamil Nadu. p. 271- 273. In A.G. Ponniah and A. Gopalakrishnan (eds.) Endemic fish diversity of Western Ghats. NBFGRNATP Publication. National Bureau of Fish Genetic Resources, Lucknow, U.P., India. 1,347 p.
2. Asahida, T., Kobayashi, T., Saitoh, K. and Nakayama, I. (1996). Tissue preservation and total DNA extraction from fish stored at ambient temperature using buffers containing high concentration of urea. Fish. Sci. 62: 727–730.
3. Chen, C. A., Odorico, D. M., Lohuis, M. T. et al. (1995). Systematic relationship within the Anthozoa (Cnidaria: Anthozoa) using the 5'-end of the 28S rDNA. Mol. Phyl. Evol. 4: 175–183.
4. Cummings, A. S. and Thorgaard, G. H. (1994). Extraction of DNA from fish blood and sperm. – Biotechniques 17: 426.
5. Dessauer, H. C., Cole, C. J. and Hafner, M. S. (1996). Collection and storage of tissues. – In: Hillis, D. M., Moritz, C. and Mable, B. K. (eds), Molecular systematics. Sinauer Ass. Inc, pp. 29–47.
6. Estoup, A., Largiader, C. R., Perrot, E. and Chourrout, D. (1996). Rapid onetube DNA extraction for reliable PCR detection of fish polymorphic markers and transgenes. Mol. Mar. Biol. Biotec. 5: 295–298.
7. Hilsdorf, A., Caneppele, D. and Krieger, J. E. (1999). Muscle biopsy technique for electrophoresis analysis of fish from the genus Brycon. Genet. Mol. Biol. 22: 547–550.
8. O‘Brien, S. J. (1994). A role for molecular genetics in biological conservation. Proc. Natl Acad. Sci. USA 91: 5748–5755.
9. Pinto, S. M., Fernandes-Mtioli, F. M. C. and Schlenz, E. (2000). DNA extraction from sea anemone (Cnidaria:Actiniaria) tissues for molecular analyses. Genet. Mol. Biol. 23: 601–604.
10. Sambrook, J. and Russell, D. W. (2001). Molecular cloning. A laboratory manual. Cold Spring Harbor Laboratory Press, New York.
11. Silas, E.G. (1953). New fishes from the Western Ghats, with notes on Puntius arulius (Jerdon), Records of the Indian Museum, 51 (1953 [1954]): 27–37.
12. Strassmann, J. E., Solis, C. R., Peters, J. M. and Queller, D. C. (1996). Strategies for finding and using highly polymorphic DNA microsatellite loci for studies of genetic relatedness and pedigrees. In: Ferraris, J. D. and Palumbi, S. R. (eds), Molecular zoology. Wiley-Liss, New York, pp. 163–180.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License