IJCRR - 4(16), August, 2012
Pages: 146-151
Date of Publication: 28-Aug-2012
Print Article
Download XML Download PDF
PROTEASE ACTIVITY OF PEPTOCOCCUS ACTIVUS
Author: Zinnat Shahina, Mohammad Towhid Hossain, Mohammad Abdul Hakim
Category: General Sciences
Abstract:Peptococcus activus are anaerobic, non motile, non spore forming gram positive cocci. The objective of this study was to investigate the availability of this bacteria and their ability to protease production. Eleven different bacterial colonies were isolated from shrimp and dry fish(shrimp).Of them, eight colonies(Z1-Z8) were isolated from shrimp and rest of the three isolated from dry fish (Z9-Z11).Among them peptococcus activus which was named Z10 before identification exhibited highest protease production at temperature 37ºC,medium pH 5.0 and 48 hours of incubation period. Maximum protease activity was achieved at 400C for 70 minutes and pH 5.0 along with 2% gelatin as substrate concentration during enzyme substrate reaction.
Keywords: Peptococcus activus , Protease, Extracellular protease, enzyme activity.
Full Text:
INTRODUCTION
Proteases represent the class of enzymes which play an important role as well as their commercial applications. Commercial proteases account for nearly 60% of the total industrial enzyme market. This proteolytic enzyme finds numerous applications in pharmaceutical, food and detergent industries [1]. Proteases of commercial importance are produced from microbial, animal and plant sources [2].Microorganisms are a great source of proteases owing to the limited space required for their cultivation and their ready susceptibility to genetic manipulation. For the production of enzymes for industrial use, isolation and characterization of new promising strain is a continuous process[3].As protease represents one of the major groups of industrial enzymes, it is worthwhile to screen microbes from new habitats for proteases with novel properties to meet the needs of rapidly growing industries[4]. The aim of this study was to isolate a bacterium capable of producing protease which was isolated from dry fish. The ability of the bacteria to grow and produce protease under different environmental conditions was studied. The microorganism was capable of producing large amount of protease when the condition varies. Effects of pH, temperature and substrate concentration of protease were evaluated.
MATERIALS AND METHODS
Microorganisms
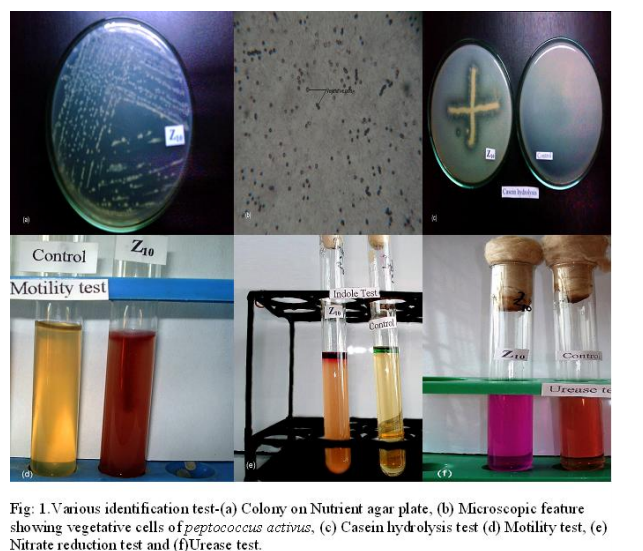
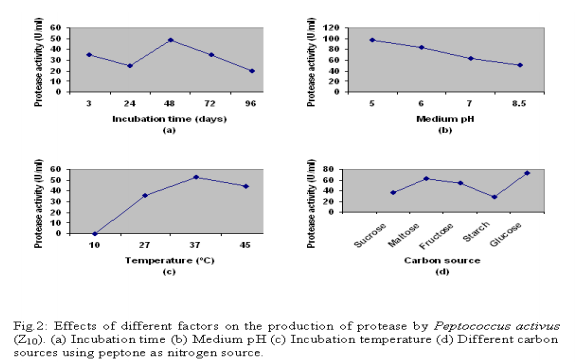
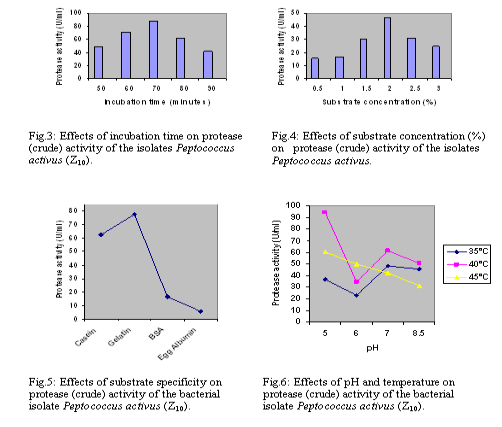
The study of protease activity by the bacterial isolate Peptococcus activus was carried out at the Department of Microbiology, University of Chittagong, starting from August 2006 to April 2007, which were isolated from dry fish(shrimp). Isolation and screening of the isolates For the isolation of proteolytic microorganisms enrichment technique was followed. Primary screening was done by three methods – Boiled egg albumin degradation, Skimmed milk casein hydrolysis and Gelatin hydrolysis. During secondary screening 3 different broth media such as (i) Peptone 2%-yeast extract 1%,dextrose 2% [5],(ii) Tryptone 1%-yeast extract 0.5%-dextrose 0.1% [6],(iii) Gelatin 1%- yeast extract-0.2%-glucose 1%-K2HPO4-0.3%- KH2PO4 0.1%-MgSO4.7H2O trace [7] broth were used. Measurement of enzyme activity Protease activity was done by the modified method of Hayashi et.al.,[8],as followed by Meyers and Ahearn [9].Enzyme solution (3ml) was incubated with 3ml of 1% casein as substrate in 3ml of citrate phosphate buffer at 350C in a water bath for 1 hour. The reaction was terminated by 20% TCA and was further incubated for 1 hour it was then filtered through Whatman no.40.One ml enzyme substrate mixture was taken into a test tube and to this 2ml of 20% Na2C03 was added, followed by 1 ml of Folin Ciocalteu Reagent was added and shaken well. After 30 minutes 6ml distilled water was added and absorbance of the solution was measured at 650 nm in a spectrophotometer. The amount of amino acid released in the reaction was calculated from a standard curve previously plotted from the known concentration of tyrosine. Enzyme activity was expressed in Unit, which was defined as the amount of enzyme that releases 1 ?g of tyrosine /ml of crude extract /hour. Optimization of culture conditions A. Effect of incubation time, temperature and medium pH To ascertain the effect of culture conditions the present study was carried out at different incubation time (24, 48, 72 and 96 hours), pH (such as 5.0, 6.0, 7.0, 8.0 and 8.5 respectively) and temperature (such as 30?2 0C, 37?2 0C, and 45?2 0C respectively).Their effect of temperature on biomass characteristic, biomass yield and protease production was recorded. B. Effect of carbon sources The production of extracellular proteases using different carbon source and the original nitrogen source (peptone) were studied in the liquid culture medium [6]. Five carbon sources (sucrose, dextrose, fructose, starch, and maltose) were added to the medium and their effect on the production of protease, extracellular protein and biomass yield were recorded. Factors affecting enzyme activity To determine the optimum condition for the crude enzyme activity several factors such as pH (ranging form 5.0 to 8.5), optimum temperature (35 to 45?2 0C), incubation time (50, 60, 70, 80 and 90 minutes ), substrate concentration (i.e. 0.5, 1.0, 1.5, 2.0, 2.5 and 3.0%) substrate specificity (using Casein, Egg albumin, BSA and Gelatin) were observed for highest enzyme activity. RESULTS AND DISCUSSION Primary screening confirmed that the strain Z10 showed greater albumin degradation, casein and gelatin hydrolysis. After primary screening this bacteria allowed to grow in three liquid media and maximum enzyme activity was found in the trypton-yeast extract-dextrose broth. On the basis of morphological (including size, shape; arrangement of the cell; presence and absence of spores; acid fast reaction, gram reaction etc), cultural, physiological and biochemical properties the strain was characterized. Then compared with the standard description of “Bergey’s Manual of Determinative Bacteriology”, 8th ed. [10]. The bacterial isolate Z10 was found to belong to the genus Peptococcus and provisionally identified as Peptococcus activus (Fig: 1.a,b,c,d, e & f). Optimization of different cultural conditions Effects of incubation period The Bacterial isolate Z10 (Peptococcus activus) showed maximum protease production 48.61 U/ml) after 48 hrs of incubation period (Fig-2.a) but highest biomass (1.020 absorbance at 600nm) was recorded after 72 hrs of incubation time. The pH of the culture filtrate were ranged from 5.05 to 5.77 . Production of maximum protease within 48-hours by bacteria was reported by Das et al. [11]. Effects of temperature The influence of incubation temperature on protease production by bacteria was studied by several workers [12-13].Among the different incubation temperatures tested for protease production 37ºC was found to be optimum for this isolate. At this temperature highest protease activity was 52.5 U/m1 and the highest biomass yield was 1.072 absorbance at 600nm (Fig-2.b). Effects of pH In the present study, maximum protease production was achieved at medium pH 5.0 (Fig-2.c). Production of protease at acidic medium pH ranged form 4.5 to 6.5 by microbes was observed by North [14]. Effect of carbon source Maximum production of protease was achieved by the isolate when gelatin was used as a nitrogen source and glucose as a carbon source (Fig-2.d). Factors involved in enzyme activity Enzyme activity depends upon the pH, temperature, incubation time, substrate concentration and many others factors. So it is necessary to find out different limiting factor for maximum activity of proteases.Thus the crude enzymes obtained were studied for the determination of different factors for maximum protease activity. Enzyme-substrate reaction incubation time The culture filtrate of bacterial isolate Z10 exhibited highest protease activity when enzyme substrate reaction mixtures were incubated at 40 ºC for 70 minutes (Fig-3) Substrate concentration The crude enzyme extract of isolate Z10 (Peptococcus activus) were allowed to react with different substrate concentrations(0.5- 3.0%) and maximum activity was recorded with 2.0% substrate concentration (fig-4). Substrate specificity Different substrate such as casein, gelatin, bovine serum albumin (BSA) and egg albumin were used to observe the efficiency of enzyme activity on different substrate and gelatin gave the highest value of activity (77.8 U/ml) with the crude of bacterial isolate Z10 (Peptococcus activus) (Fig-5). Temperature and pH Temperature and pH are also most important limiting factors, which markedly influenced enzyme activity. Maximum protease activity of crude enzyme extract of bacterial isolate Z10 was recorded at temperature 40ºC and pH 5.0 (Fig:6).similar temperature was also observed by Lee [15].
CONCLUSION
The inability of the plant and animal proteases to meet current world demands has led to an increased interest in microbial proteases. Proteases from microbial sources are preferred to the enzymes from plant and animal sources since they possess almost all the characteristics desired for their biotechnological applications. Despite the extensive research on several aspects of proteases, there is a paucity of knowledge about the roles that govern the diverse specificity of these enzymes. Deciphering these secrets would enable us to exploit proteases for their application in biotechnology. Further investigation on this bacterial (peptococcus activus) species can possibly reveal it’s potentiality as a source of protease for any biotechnological approach.
ACKNOWLEDGEMENTS
Authors acknowledge the immense help received from the scholars whose articles are cited and included in references of their manuscript. The authors are also grateful to authors/editors/ publishers of all those articles, journals and books from where the literature for this article has been reviewed and discussed.
References:
1. Moon, S. H. and S.J. parulekar. 1991. A parametric study of protease production in batch and fed-batch cultures of Bacillus firmus. Biotechnol. Bioengg., 37: 467-483.
2. Patel, P.R., 1985.Enzymes iaolation and purification. In: Biotechnology: Applications and Research. Technomic Publishing Co. Inc., USA, pp: 534-564.
3. Kumar., A. Sachdev., A. Balasubramanyam., SD. Saxena., AK. And Lata, 2002. Optimization of conditions for production of Neutral and Alkaline protease from species of Bacillus and Pseudomonas. Ind. J. Microbiol.42:233-236.
4. Kumar, D., Savitri, N. Thakur, R. Verma and T. C. Bhalla, 2008.Microbial proteases and application as laundry detergent additives. Res. J. Microbial., 3: 661-672.
5. Amar, J.S., and Halvorson, H.O. 1975. Proteinase activities of Saccharomyces cerevisiae during sporulation. Journal of Bacteriol. 124(2): 863-869.
6. Matta,H.,V.Punj and S.S.Kanwar,1997.An immuno-dot assay for detection of thermostable protease from Pseudomonus sp.AFT-36 of dairy origin.Appl.Microbiol.,25:300-302.
7. Shalinisen and Satyanarayana, T. 1993. Optimization of alkaline protease production by thermophilic Bacillus licheniformis S-40. Indi J. Microbiol. 33(1): 43-47.
8. 8.Hayashi,L.D.,D.Fukushima and K.Mogi,1967.Agr.Biol.chem.., Tokyo,31:1237-1241.Cited from Physiology and Parasitism.Edt.By G.P.Agarwal and K.S.Bilgrami.Today and tomorrow’s printers and publishers,New Delhi,India,pp:194.
9. Meyers,S.P. and D.G.Ahearn, 1977. Mycologia,69:649-651. Cited from Physiology and Parasitism.Edt.By G.P.Agarwal and K.S.Bilgrami.Today and tomorrow’s printers and publishers,New Delhi,India,pp:195.
10. Buchanan,R.E. and Gibbons,N.E.(8th eds),1974.The William and Wilkons Co.Baltimore.
11. Das,F.,Hossain,MD.T.and Anwar,M.N.2005.Protease production by Bacillus Sphericus.TheChittagong univ.j.Sci,29(1):71-76.
12. Al-Saleh, A.A. and Zahran, A.S. 1997. Protease production by Pseudomonas fleorescens CM12 isolated from raw camel milk. Egypt. J. Dairy Sci. 25 (2): 327-336.
13. Sharmin, S., Hossain, M.T. and Anwar,M.N.2005. Isolation and characterization of a protease producing Bacteria Bacillus amovivorus and optimization of some factors of culture conditions for protease production, Journal of Biological Sciences 5(3):358-362.
14. Lee W., Young-Jecho, Gyu-Mok S. and Cheong Choi. 1992. Characteristic and action pattern of alkaline protease produced from Bacillus sp. CW-1121. Korean Biochem J. 24(6): 537-542.

|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License