IJCRR - 4(24), December, 2012
Pages: 07-16
Print Article
Download XML Download PDF
ANTIFUNGAL ANTIBIOTIC PRODUCTION BY STREPTOMYCES SP. ISOLATED FROM SOIL
Author: Shipra Singh, Anita Rawat, Deepak Chand Sharma
Category: General Sciences
Abstract:19 Actinomycetes isolate were obtained from soils samples collected from various ecological niches of Western Uttar Pradesh (Meerut). They were selected for their antimicrobial producing capabilities against some selected microbial strains including fungus (Aspergillus sp.), Yeast (Candida sp.), gram-stain-positive bacteria (Bacillus sp.) and gram-stain-negative bacteria (Burkholderia sp.). Out of 19 isolates, 12 (63%) were found to have antimicrobial activity, among them 53% and 26% isolates were found active against gram-stain-positive and gram-stain-negative bacteria, respectively. Antifungal activity was recorded in 42% isolates, among them 32% were active against Aspergillus sp. and 11% were found active against Candida sp. Only one isolate (SS12) was found to produce broad spectrum antibiotic effective against all test microorganisms. 16S rDNA sequencing and BIOLOG analysis of SS12 suggested that it may be a novel species of genus Streptomyces.
Keywords: Antifungal antibiotic, BIOLOG, 16S rDNA, Streptomyces, Actinomycetes
Full Text:
INTRODUCTION
Actinomycetes, group of heterotrophic, hyphae forming, gram-stain-positive bacteria identified as one of the major group of soil population (Kuster, 1964). These prokaryotes have diverse metabolic requirements and have been explored and exploited for ages in search of various primary and secondary metabolites, such as antibiotic, enzymes and immune modulators (Moncheva et al., 2002). Approximately 70-80% of the antibiotic market is attributed to Actinomycetes; among them more than 50% is contributed by the genera Streptomyces and Micromonospora (Pandey et al., 2004). According to World Health Organization over prescription, excessive and improper use of antibiotics has resulted in the development of Multiple Drug Resistant (MDR) strains of pathogenic microbes. The prevalence of antimicrobial resistance among key microbial pathogens is increasing at an alarming rate worldwide (Singer et al., 2003). This strike back of pathogens is increasing globally and may render the current antimicrobial agents insufficient to control at least some microbial infections (Walsh et al., 2011). This makes the search of novel antimicrobial agents with clinical importance more significant to counter drug resistant microbes. In the present study we have focused on isolation of Actinomycetes from terrestrial soils in ecological niches and check their potential to produce antimicrobials.
MATERIAL AND METHODS
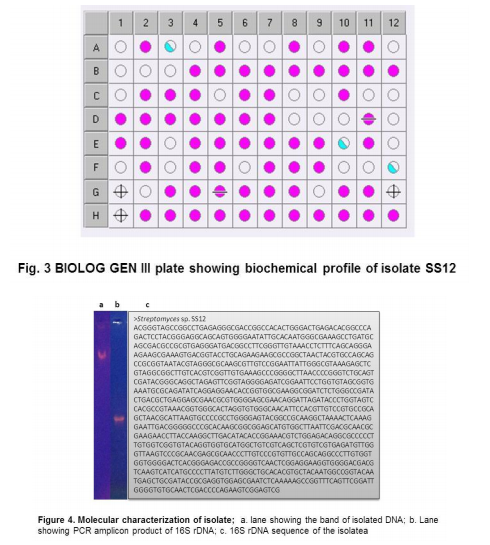
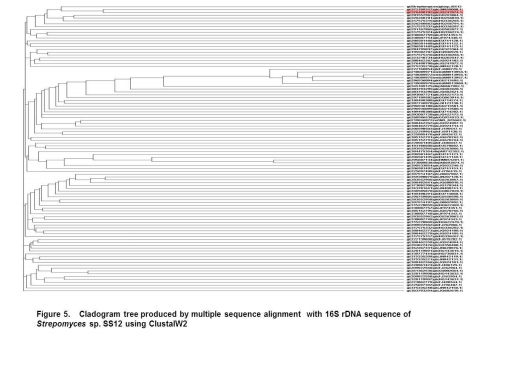
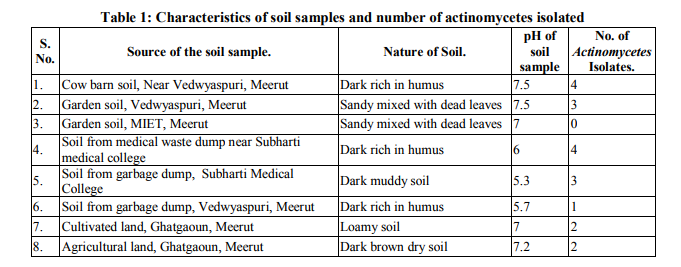
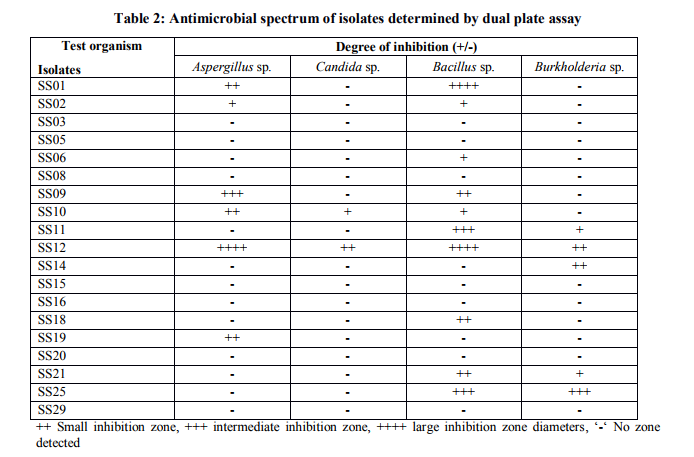
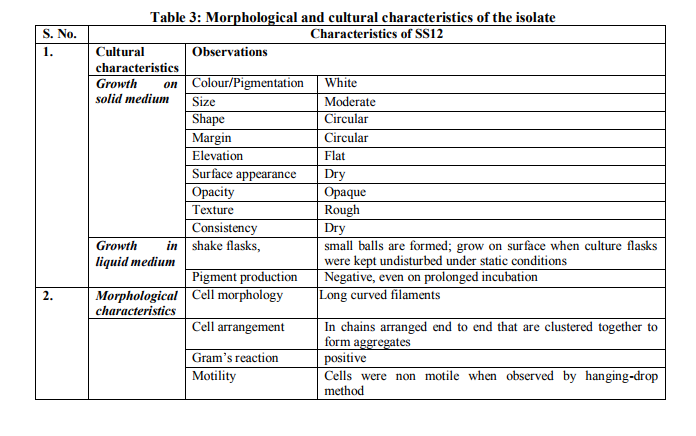
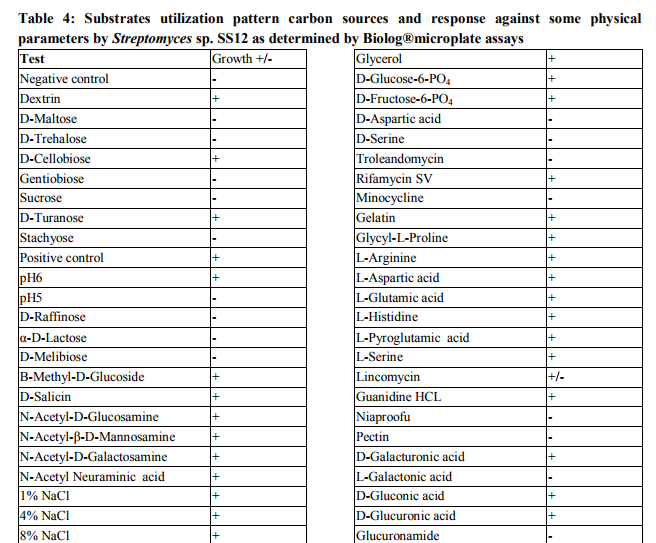
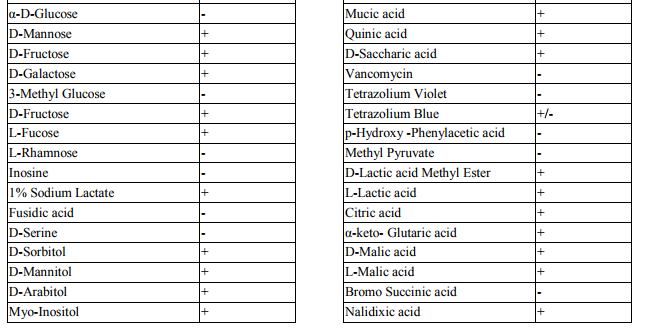
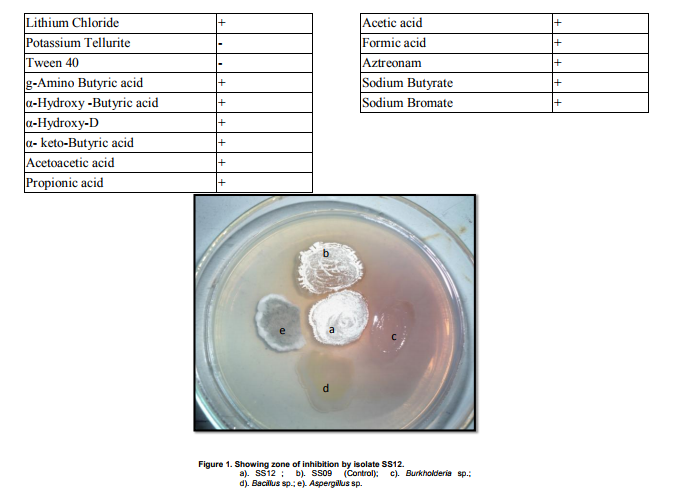
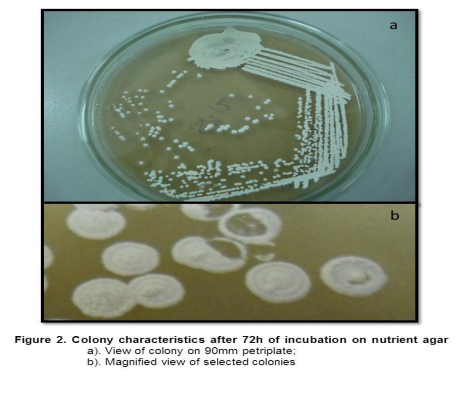
Collection of soil samples Several diverse habitats were selected for the isolation of Actinomycetes. These habitats include cow barns, garden soil, medical waste dumps, garbage dumps etc. Soil samples collected from at least 4 different places at each site. The samples were taken from a depth of 10- 15 cm after removing top soil (Ogunmwonyi et al., 2008), placed in polyethylene bags, closed tightly and stored in a refrigerator at 4°C and processed with in 24h. Test microorganism The test microorganisms used to evaluate the antimicrobial property of isolates were obtained from Subharti medical college, Meerut. The strains represent the various groups of microbes like Aspergillus sp. (Mycellial form of fungi), Candida sp. (Budding form of fungi), Bacillus sp. (gram-stain-positive) and Burkholderia sp. (gram-stain-negative). Isolation of Actinomycetes 1g of soil sample was mixed aseptically in 100mL of sterilized normal saline, and maintained at 50-55 ºC for 1h followed by spreading on petri plates containing Actinomycetes Isolation Agar (AIA) and plates were incubation at 28°C for 10 days (El-Nakeeb and Lechevalier, 1963; Kuster and Williams, 1964). The colonies showing typical morphology (Williams and Cross, 1971) were purified and stored at 4°C in agar slants and as glycerol stock at -20°C. The pretreatment with heat enhances the population of Actinomycetes in soil sample. Screening of Actinomycetes for antimicrobial activity Dual plate assay method The center of PDA plates was point inoculated with test fungi and antagonist bacterial cultures were point-inoculated at the periphery of the plate. Each plate was incubated at 28?C, for 72- 96 hours in an inverted position (Huang et al., 1976). The zone of inhibition of the fungus around each isolate was measured. Identification of the potent Actinomycetes isolate The isolated cultures (Actinomycetes) were classified on the basis of its phenotypic, morphological and biochemical characters. Morphological and microscopic characterization Morphological and cultural characters of the selected actinomycetes strains were studied by inoculating the selected strain into sterile ISP media. The media were sterilized and poured into sterile Petri dish. After solidification of the media, the culture of the selected strain was streaked on the media surface aseptically and incubated at 28 °C for 7 days. Morphological properties such as colony characteristics, type of areal hyphae, aerial mass colour, growth of vegetative hyphae, reverse side pigments, melanoid pigments, fragmentation pattern, spore formation and spore chain morphology were observed (Shirling and Gottileb, 1966).A smear of the selected strain was prepared on a clean glass slide and after performing Gram’s staining was examined under oil immersion (100 X) (Williams, 1993). Biochemical characterization of isolates The isolate was biochemically characterized by sending the cultures to BTK biosciences, New Delhi for Biolog® system analysis (Biolog, USA). Molecular Identification of the strain SS12 DNA isolation of SS12 10 mL of nutrient broth was inoculated with the bacterial isolate and incubated at 30ºC in shaker at 250 rpm. After 12 h, 100µL of glycine (0.1%) solution was added to broth culture and incubated further for 6 h. Genomic DNA was extracted accoeding to Bazzicalupo and Fani (1994). Extracted genomic DNA was electrophoresed on 0.8% agarose gel in TRIS-acetate-EDTA (TAE) buffer (1X) at 80 V for 60 min. Ethidium bromide (0.5 g mL-1 ) was added at the time of gel casting. After the run, gel was visualized under UV transilluminator. Amplification of 16S rDNA 16S rDNA regions of bacterial isolates were amplified from bacterial genomic DNA by polymerase chain reaction, using eubacterial universal primers: Gm3f 5 AGA GTT TGA TCM TGG 3 (8 to 23) Gm4r 5 TAC CTT GTT ACG ACT T 3 (1492 to 1507) Reaction was set in 50µL volume with sterile triple distilled water, Taq DNA Pol. and MgCl2 15 mM (1X), dNTP mix (10 mM, 0.25 mM), Primer Gm3f (0.25 M), Primer Gm4r(0.25 M), Taq DNA polymerase (1.0 U),Target DNA (template), 20-100ng. The amplification cycle consisted of initial denaturation step at 95 °C for 7 min followed by 25 consecutive cycles of 60 sec at 94 °C, 60 sec at 51 °C, 60 sec at 72 °C and three touchdown cycles were successively performed at 54, 53 and 52 °C followed by final extension at 72 °C for 10 min. Positive and negative controls were invariably maintained. The PCR product was run on 0.8% agarose gel and visualized under UV. Sequencing of isolate SS12 The 16S rDNA sequence of the bacterial isolates was determined by dideoxy chain termination method by sending the samples to Chromus India Pvt. Ltd. (Banglore, India). Analysis of sequence data and identification of the bacterium Blast database of the National Centre for Biotechnology information (NCBI) was used to compare the sequences of isolates with known 16S sequences in the existing database. The results obtained were also confirmed by comparing with Ribosomal Database Project. Sequences were aligned through multiple sequence alignment, ClustalW programme, from the European Bioinformatics Institute (EBI) accessible on net (http://www.ebi.ac.uk/clustalW) and phylogram was constructed to understand evolutionary relatedness using MEGA 3.0 (Kumar et al. 2001). Nucleotide sequence accession numbers The 16S rDNA nucleotide sequence obtained in the study was submitted to the GenBank database. RESULT AND DISCUSSION Screening of soil sample Soil samples were collected from eight different locations in Meerut. Emphasis was given to collect soil from places where industrial wastes and household garbage were being dumped. Total 31 actinomycetes strains with varying morphological characteristics were picked up and their pure culture was maintained at 4°C for further studies. Pertinent details of the soil samples and the actinomycetes are shown in Table 1. Soil is known as reach source of Actinomycetes having antifungal activity against plant fungal-pathogens, 110 isolates were screened by Aghighi et al. (2004), from which 14 isolates were found active against various fungal isolates. Screening of isolates for antimicrobial activity Dual plate assay The antimicrobial spectrum of the actinomycetes isolates was confirmed by growing the actinomycetes isolate with the test cultures (Aspergillus sp., Candida sp., Bacillus sp. and Burkholderia sp.). Out of 19 isolates 12 (63%) showed antimicrobial activity against at least one test organism. 53% isolates were found active against at least one gram-stain-positive bacteria and 26% active against at least one gram-stainnegative bacteria and only 32% isolates were active against the fungal test microorganism. Onlyten (53%) isolates were effective against Bacillus sp., five (26%) against Burkholderia sp., two (11%) against Candida sp. and only six (32%) were found to be active against Aspergillus sp. Only one isolate was found effective against all the microorganisms tested (Fig. 1 and Table 2). In a similar study (Oskay, 2004), 50 isolates of actinomycetes were isolated from 10 farming soil samples collected in Manisa Province, Turkey. Approximately 34% of the isolates produced broad and narrow spectrum antibiotics, 16% of the isolates produced antibacterial substance that were active against only gram-stain-positive bacteria 6% of the isolates were active against gram-stain-negative bacteria and 12 % of the isolates were active against both gram-stain-positive and gram-stain-negative bacteria. SS12 was selected for further study due to larger zone size and broad spectrum of activity. Morphological and cultural characteristics of strain SS12 Microscopic observation (1000X magnification) after Gram’s staining revealed that SS12 is a Gram-stain-positive and rod-shaped microorganism (Hucker and Conn, 1923). Other morphological characteristics such as colony characteristics, type of areal hyphae, aerial mass colour, and growth of vegetative hyphae, reverse side pigments, melanoid Pigments, fragmentation pattern and spore formation are detailed in Table 3. They indicate that strains SS12 belongs to the genus, Actinomycetes (Fig. 2) (Shirling and Gottlieb, 1966). Biochemical characterization of isolates Biochemical properties of isolate did not matched with the available data of BIOLOG. The suggested match didn’t belong to actinomycetes; two major possibilities were worked out including novel species or contamination. Second possibility was rejected by the fact that positive and negative control was normal (Fig 3, Table 4). Identification of actinomycetes by biochemical properties are of prime interest (Moncheva et al., 2002) Analysis of sequencing data and identification of the isolates The band of genomic DNA was observed near well and PCR amplicon had a molecular weight of 1.5kb (Fig 4b). The 16S rDNA of the isolate was sequenced, and the sequence of the isolate is shown in Fig 4c. The sequence of 16S rDNA obtained was searched for homogenous sequences in the GenBank database. The results obtained were confirmed by NCBI. This bacterial strain was identified as the member of the genus Streptomyces (BankIt1564851). The isolate did not showed similarity with known isolates of the database. As shown by similarity tree constructed by ClustalW2, and designated as Streptomyces sp. SS12 (Fig. 5). CONCLUSION Nineteen isolates from terrestrial soil of ecologically stressed niches were isolated and their antimicrobial activity against bacteria and fungi was studied. One isolate was found to be the producer of broad spectrum antibiotic and its morphological, cultural and molecular identification indicate that the isolate belongs to genus Streptomyces. Further studies to optimize the culture parameter for economic production of antifungal antibiotic resulted in the enhancement of production level.
References:
1. Aghighi S, Bonjar GHS, Rawashdeh R, Batayneh S and Saadoun I. First report of antifungal spectra of activity of Iranian Actinomycetes strains against Alternaria solani, Alternaria alternate, Fusarium solani, Phytophthora megasperma, Verticillium dahlia and Saccharomyces cerevisiae. Asian J Plant Sci 2004; 3(4): 463-471.
2. Bazzicalupo M, and Fani R. The use of RAPD for generating specific DNA probes for microorganisms, in Methods in Molecular Biology (Clapp, J. P., ed.), Humana, Totowa, NJ, 1995; 155–175.
3. El-Nakeeb MA and HA Lechevalier. Selective isolation of aerobic actinomycetes. Appl Microbiol 1963; 11: 75-77.
4. Huang HC and Hoes JA. Penetration and infection of Sclerotinia sclerotiorum by Coniothyrium minitans. Can J Bot 1976; 54: 406-410.
5. Hucker GJ and Conn HJ. Methods of Gram staining. Technical Bulletin of the New York State, Agric Exp Station, 1923; p. 93.
6. Kumar, Tamura SK, Jakobsen IB, and Nei M. MEGA2: molecular evolutionary genetics analysis software. Bioinformatics 2001; 17:1244-1245.
7. Kuster E, and Williams ST. Selection of media for isolation of streptomycetes. Nature (London) 1964; 202:928- 929.
8. Moncheva P, Tishkov S, Dimitrova N, Chipeva V, Antonova-Nikolova S and Bogatzevska N. Characteristics of soil Actinomycetes from Antarctica. In: Journal of culture collections 2002;3: 03-14.
9. Moncheva P, Tishkov S, Dimitrova N, Chipeva V, Stefka AN, and Bogatzevska N. Characteristics of soil actinomycetes from Antarctica, Journal of culture collections 2002; 3, 3-14.
10. Ogunmwonyi IN, Igbinosa OE, Aiyegoro OA and Odjadjare EE,. Microbial analysis of different top soil samples of selected site in ObafemiAwolowo University, Nigeria Scientific Research and Essay 2008; 3 (3): 120-124.
11. Oskay MA, Usame T, and Cem A,. Antibacterial activity of some actinomycetes isolated form farming soils of Turkey. Afr J Biotechnol 2004; 3(9): 441-446.
12. Pandey B, Ghimirel P, Prasad V, Thomas M, Chan Y, and Ozanick S. Studies of the antimicrobial activity of the actinomycetes isolated from the Khumby region of Nepal. Department of Bacteriology, University of Wisconsin— Madison, Madison, Wisconsin. 2004.
13. Singer RS, Finch R, Wegener HC, Bywater R, Walters J and Lipsitch M. Antibiotic resistance – the interplay between antibiotic use in animals and human beings. Lancet Infect Dis 2003; 3:47-51.
14. Walsh TR, Janis W, David ML and Mark AT. "Dissemination of NDM-1 positive bacteria in the New Delhi environment and its implications for human health: an environmental point prevalence study". The Lancet Infectious Diseases 2011; 11 (5): 355–362.
15. Williams ST and Cross T. Actinomycetes. Applied microbiology. 4, Academic Press; 1971.
16. Williams ST, Locci R, Beswick A, Kurtboke DI, Kuznetsov VD, Lemonnior FJ and Long PF. Detection and identification of novel actinomycetes. Res Microbiol 1993; 144(8):653-656.


|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License