IJCRR - 5(1), January, 2013
Pages: 44-48
Print Article
Download XML Download PDF
INDUCIBLE CLINDAMYCIN RESISTANCE AMONG CLINICAL ISOLATES OF METHICILLIN RESISTANT STAPHYLOCOCCUS AUREUS
Author: Nilima R. Patil, Ulhas S. Mali, Sunanda A. Kulkarni, M. V. Ghorpade, Poorva P. Bhave (Sule)
Category: Healthcare
Abstract:The resistance to antimicrobial agents among Methicillin resistant Staphylococcus aureus (MRSA) is an increasing problem. Clindamycin is commonly used for the treatment of serious soft tissue infections produced by Staphylococcus aureus and treatment failures are reported during therapy. Routine antibiotic sensitivity test for Clindamycin susceptibility may fail to identify iMLSB (inducible MLSB) strains of Staphylococcus aureus due to erm gene. Present study was carried out to find out the percentage of inducible Clindamycin resistance in MRSA in our hospital using D-test. 250 isolates of Staphylococcus aureus from various clinical samples were confirmed by standard protocol and included in this study. MRSA were detected by using Oxacillin (1ug) and Cephoxitin (30ug) disc diffusion method. Results were interpreted according to CLSI criteria. Antibiotic sensitivity to routine antimicrobial agents was carried out using Kirby Bauers disc diffusion method. D-test was performed on all Erythromycin resistant Staphylococcus aureus to detect inducible Clindamycin resistance and constitutive Clindamycin resistance. Out of 250 isolates of Staphylococcus aureus, 90(36%) were found to be MRSA. Among these, 46(51.11%) were Erythromycin resistant. 17(36.95%) isolates showed inducible Clindamycin resistance, 4(8.69%) showed constitutive Clindamycin resistance and 25(54.34%) showed MS phenotype in MRSA. Conclusion \? We therefore conclude that it is necessary to perform D- test for detection of inducible Clindamycin resistance among Staphylococcus aureus in routine antibiotic sensitivity test. Therapeutic failures may thus be avoided.
Keywords: Staphylococcus aureus, inducible MLSB, constitutive Clindamycin resistance, D-test, MRSA.
Full Text:
INTRODUCTION
Staphylococcus aureus is one of the most common human pathogens which cause wide range of infections. The emergence of resistance to antimicrobial agents among Staphylococci is an increasing problem. Emergence of methicillin resistance in Staphylococcus aureus has resulted in therapeutic alternatives to treat Staphylococcal infections. The Macrolides (Erythromycin, Clarithromycin) and lincosamides (Clindamycin, Lincomycin) serve as one such alternative. Macrolides, Lincosamides and Streptogramin B are structurally unrelated but microbiologically having same mode of action (1). Clindamycin is commonly used for the treatment of serious Staphylococcal infection due to its excellent pharmacokinetic properties but sometimes treatment failures were reported during therapy (2, 3). However widespread use of Macrolideslincosamides-streptogramin B (MLSB) antibiotics has led to an increased resistance to these antibiotics by Staphylococcal strains (4). The well established mechanism for such resistance is target site modification mediated by erm gene which can be expressed either constitutively or inducibely. Erythromycin & Clindamycin discs are commonly used while doing antibiotic sensitivity test for Staphylococcus aureus strains. Inducible resistance to Clindamycin is difficult to detect in routine antibiotic sensitivity test as organisms are resistant to Erythromycin and sensitive to Clindamycin in vitro when not placed adjacent to each other. In such cases, in vivo therapy with Clindamycin may select constitutive erm mutants leading to clinical therapeutic failure. Another mechanism of resistance to Macrolides by MRSA is efflux of antibiotic. Staphylococcal isolates appear Erythromycin resistant and Clindamycin sensitive in both in vivo and in vitro and the strain does not typically become Clindamycin resistant during therapy (5). The present study was carried out to find out inducible Clindamycin resistance in MRSA & interpretation of susceptibility test to guide therapy.
MATERIAL AND METHODS The present study was conducted from November-2010 to November -2011. A total 250 isolates of Staphylococcus aureus were isolated from various clinical specimens like pus, wound swabs, blood and various aspirations received in the department of Microbiology, B.V.D.U.M.C & H. Sangli. All the Staphylococcus species were identified by conventional microbiological methods including colony morphology, gram stain, slide coagulation test, tube coagulation test and mannitol fermentation test, and then subjected to susceptibility testing by Kirby – Bauers disc diffusion method on Muller Hinton agar plate using routine antibiotic discs as per CLSI guidelines. Methicillin resistance was detected by using Oxacillin (1-ug) and Cephoxitin (30-ug) on Muller-Hington agar supplemented with 2% NaCl followed by incubation at 35oC. (6) The isolates which were found to be Erythromycin resistant were examined for inducible Clindamycin resistance [iMLSB] by using double disc approximation test (D test) as per CLSI guidelines. Erythromycin (15-ug) disc was placed at a distance of 15 mm (edge to edge) from Clindamycin (2-ug) disc on MullerHington agar plate. After overnight incubation at 37oC, flattening of zone (D shaped) around Clindamycin in the area between the two discs indicated inducible Clindamycin resistance. Strains that do not form inhibition zone around Clindamycin and Erythromycin discs are accepted as constitutive to MLSB phenotype (cMLSB). (6, 7) Figure No. 1, 2, and 3 show iMLSB, cMLSB and MS phenotype.
RESULT Specimen wise distribution of 250 samples is shown in table No.1. Out of the 250 Staphylococcus aureus isolates, 90 were found to be MRSA. Among these, 46 (51.11%) were Erythromycin resistant. D-test was performed for these isolates, 17 (36.95%) isolates showed inducible MLSB phenotype (D-test positive), 4 (8.69%) showed constitutive MLSB phenotype and 25 (54. 34%) showed D- test negative suggesting MS phenotype. (Table No. 2)
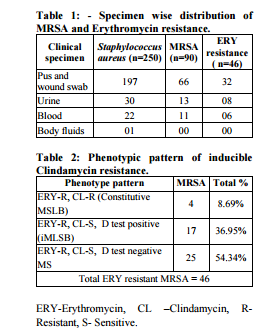
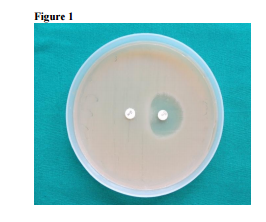
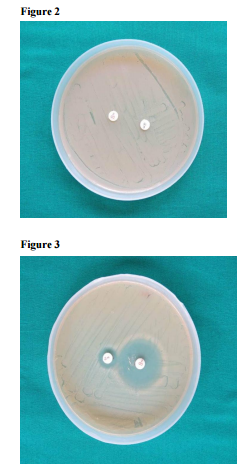
DISCUSSION MRSA strains which cause serious infections may give serious problems in treatment. Prevalence of MRSA has been reported to vary between 9 and 54.6% in hospital based infections (8, 9, 10). Characteristically, reports from different regions have shown different patterns of resistance. Some reports show high prevalence while others show low prevalence. Prevalence of MRSA in our study was 36%. A study from south Maharashtra had reported that more than 90% isolates are resistant to Penicillin, Ampicillin, Erythromycin and Gentamycin where as only 39.1% were resistant to Methicillin (14). Kandle et al showed 32.8% MRSA (15) and study from Nepal had shown 31.60% MRSA (16). Our results correlate with all of them.
The determination of antimicrobial susceptibility of a clinical isolate is often crucial for optimal antimicrobial therapy of infected patients. Resistance to macrolide, lincosamides, and streptogramin B [MLSB] antibiotics most only results from acquisition of Erythromycin resistant methylase gene (erm gene) which encodes enzymes that methylate the 23S rRNA. In our study iMLSB was found in 36.95% MRSA which is less than P. Sreenivasulu Reddy et al who observed it in 46.2% cases in MRSA (11) but is more than Yilmaz et al who found it in 24.4% and Gadepalli et al who found in 30% of MRSA cases (12) (4).
Constitutive resistance in our study is seen in 8.69% of MRSA isolates which is in accordance with V. Deotale-2010 study (7.3%) (17). 54.34% of our isolates of MRSA were sensitive to Clindamycin, against which it would be safe & appropriate to use Clindamycin or other Macrolides. It correlates with previous studies who have reported 57% of susceptibility towards Clindamycin among MRSA stains (13). Above studies showed that prevalence of inducible resistant isolates may differ from place to place.
CONCLUSION Our observations suggest that, D-test should be mandatory for all microbiological laboratories before reporting Clindamycin susceptibility as Clindamycin is not a suitable drug for D- test positive isolates while it can definitely prove to be a drug of choice in case of D - test negative isolates. Therefore, regular surveillance of antimicrobial susceptibility pattern of MRSA, determination of phenotypic pattern of inducible Clindamycin resistance and formulation of a definite antibiotic policy may be helpful in reducing the burden of MRSA infections and failures in Clindamycin treatment in the hospital.
ACKNOWLEDGEMENT Authors acknowledge the immense help received from the scholars whose articles are cited and included in references of this manuscript. The authors are also grateful to authors / editors / publishers of all those articles, journals and books from where the literature for this article has been reviewed and discussed.
References:
1. Fiebelkorn KR, Crawford SA, McElmeel ML, et al. Practical disc diffusion method for detection of inducible Clindamycin resistance in Staphylococcus aureus and coagulase negative Staphylococcus. J Clin Micro-biol 2003; 41: 4740-4744.
2. George K siberry GK, Tekle T, Carroll K, Dick Failure of Clindamycin treatment of methicillin resistant Staphylococcus aureus expressing inducible Clindamycin resistance in vitro. Clin Infect Dis 2003; 37:1257–60.
3. Lewis JS 2nd, Jorgensen JH. Inducible Clindamycin resistance in Staphylococci: should clinicians and microbiologists be concerned? Clin Infect Dis 2005; 40: 280-5.
4. Gadepalli R, Dhawan B, Mohanty S, Kapil A, Das BK, Choudhary R. inducible Clindamycin resistance in clinical isolates of Staphylococcus aureus. Indian J med Res 2006; 123: 571-3.
5. Steward CD, Raney PM, Morell AK, et al. Testing for induction of Clindamycin resistance in erythromycin resistant isolates of Staphylococcus aureus. J Clin Micro-biol 2005; 43: 1716-17121.
6. Clinical and laboratory standards institute. Performance standards for antimicrobial susceptibility testing; seventeenth informational supplement. Clinical Laboratory Standards Institute. 2007;Vol.27 (No.1)
7. Laclercq R. Mechanisms of resistance to Macrolides and lincosamides: Nature of resistance elements and their clinical implications. Clin Infect Dis. 2002; 34:482– 492.
8. Manian FA, Meyer PL, Setzer J, Senkel D. Surgical site infections associated with methicillin-resistant Staphylococcus aureus: do postoperative factors play a role? Clin. Infect. Dis., 2003, 37: 863-868.
9. Karadenizli A. Hastanelerde metisilin dirençli Staphylococcus aureus (MRSA) kontrol politikalar? ve MRSA. Hastane ?nfeksiyonlar? Dergisi, 2002, 6: 12-18.
10. Orret FA, Land M. Methicillin Resistant Staphylococcus aureus Prevalence: Current susceptibility pattern in Trinidad. BMC. Infect. Dis. 2006, 5;6:83-85.
11. P. Sreenivasulu Reddy, R. Suresh, Phenotypic detection of Clindamycin resistance among the clinical isolates of Staphylococcus aureus by using lower limit of inter disk Space.J. Microbiol. Biotech. Res., 2012, 2 (2): 258-64.
12. Yilmaz G, Aydin K, Iskender S, Caylan R, Koksal I. Detection and prevalence of inducible clindamycin resistance in staphylococci. J Med Microbiol. 2007; 56:342–5.
13. Delialioglu N, Aslan G, Ozturk C, Baki V, Sen S, Emekdas G. Inducible Clindamycin resistance in staphylococci isolated from clinical samples. Jpn J Infect Dis. 2005;58:104–6.
14. Mehta et al control of methicillin resistant Staphylococcus aureus in the tertiary care center. A five year study. J. Med. Microbiolol 1998; 16:31-4
15. Kandle S.K. et al. Bacteriophage typing and antibiotic sensitivity pattern of Staphylococcus aureus from clinical specimens in and around Solapur (SouthMaharastra). J Commun Dis 2003; 35:17- 23.
16. Pal N, Sharma B, Sharma R, Vyas L (2010). Detection of inducible Clindamycin resistance among Staphylococcal isolates from different clinical specimens in western India. J. Postgrad. Med., 56(3):182-185.
17. Deotale V, Mendiratta DK, Raut U, Narang P. Inducible Clindamycin resistance in Staphylococcus aureus isolated from clinical samples. Indian J Med Microbiol. 2010;28( 2):124–126.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License