IJCRR - 6(3), February, 2014
Pages: 16-21
Print Article
Download XML Download PDF
HIGH LEVEL AMINOGLYCOSIDE RESISTANT ENTEROCOCCUS SPECIES : A STUDY
Author: Niharika Lall, Silpi Basak
Category: Healthcare
Abstract:Background and Objectives: The emergence of Enterococcus species as a causative agent of Health care Associated Infections, poses therapeutic challenge to clinicians. Enterococci are well equipped with intrinsic resistance to multiple antibiotics. Acquired resistance to commonly used antibiotics such as Penicillin, Aminoglycoside and Vancomycin have made the situation even worse. Detection of High Level Aminoglycoside Resistance (HLAR) in Enterococcus species can predict the loss of synergy between cell wall active antimicrobial agents and Aminoglycosides. Hence, the present study was undertaken to detect the incidence of High level Aminoglycoside Resistant (HLAR) Enterococcus species isolated in a rural hospital and to study their antibiotic susceptibility profile. . Method: HLAR in Enterococcus species was detected by disk diffusion test using High level Streptomycin (HLS - 300 \?g) disk and High level Gentamicin (HLG - 120 \?g) disk and Agar dilution method as per CLSI guidelines. High Level Gentamicin Resistance (HLGR) was also confirmed by HiMedia Ezy MIC Strip. Antibiotic sensitivity test was done by Kirby Bauer Disc diffusion method as per CLSI guidelines. Result: Out of total 190 Enterococcus species isolated from different clinical samples, High Level Aminoglycoside Resistance was detected in 115 (60.5%) Enterococcus strains. Amongst 115 HLAR Enterococcus strains, 59 (51.4%) were E. faecalis and 56 (48.6%) were E. faecium. Conclusion: We hereby conclude that Enterococcus strains, isolated from different clinical specimens must be screened routinely for HLAR (High Level Aminoglycoside Resistance) by all Clinical Microbiology Laboratories to improve the therapeutic outcome.
Keywords: Enterococci, High Level Aminoglycoside Resistant (HLAR), Multi-drug resistant.
Full Text:
INTRODUCTION
Enterococci are Gram positive bacteria that normally inhabit the gastrointestinal tract of many animals including humans. However, when they colonize the sites where they are not normally found, they can become pathogens1 . Enterococci are now the second most common cause of nosocomial urinary tract and wound infections and third most common cause of nosocomial bacteremias2 . Serious enterococcal infections (e.g., bacteremia and endocarditis) require treatment with a bactericidal combination of antibiotics that should include penicillin group of drugs (e.g., Ampicillin or Penicillin G) to which the Enterococcus isolate is susceptible and an Aminoglycoside (e.g., Gentamicin or Streptomycin) to which the Enterococcus isolate does not exhibit high-level resistance3 . High-level Aminoglycoside resistance (HLAR) among Enterococci is increasingly being reported worldwide. The presence of HLAR is predictive of the loss of synergy between a cell-wall-active agent (e.g. Penicillin, Ampicillin or Vancomycin) and an Aminoglycoside, which makes the treatment of serious enterococcal infections difficult4 . Hence, the present study was undertaken to detect the incidence of High-level aminoglycoside resistant (HLAR) Enterococcus species isolated from different clinical samples and their antibiotic susceptibility profile.
MATERIAL AND METHODS
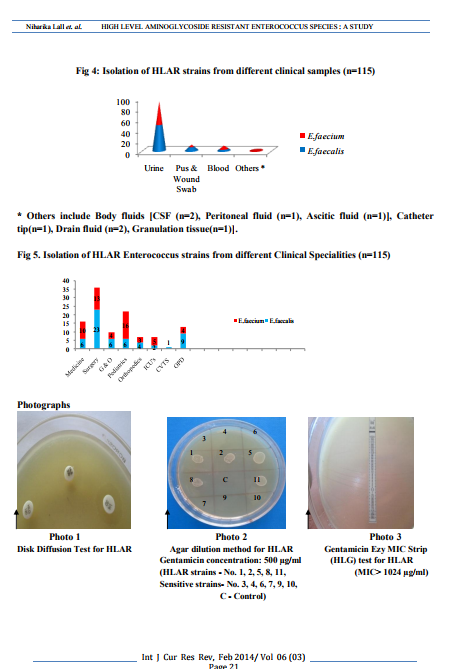
A total number of 190 Enterococci were isolated from various specimens like urine, blood, pus and wound swab, body fluids, etc from July 2011 to August 2013. The specimens were collected from patients attending Indoor and Outdoor Patients Department of our Hospital and sent to Department of Microbiology. Our hospital is a tertiary care hospital in a rural set up. All Enterococcus species were identified by conventional methods2 . The speciation of Enterococcus species was done by scheme proposed by Facklam and Collins5 . Antibiotic susceptibility test was done for all 190 strains by Kirby- Bauer disk diffusion method, according to CLSI guidelines6 . Antibiotics like Ampicillin(10µg), Linezolid (15µg),Vancomycin (30µg), Erythromycin (15µg), Tetracycline (30µg), Quinupristin-dalfopristin (15µg) were used. Additionally, for urine samples only, Nitrofurantoin disc (300µg) was used. HLAR (High Level Aminoglycoside Resistance) was detected by disk diffusion test using High level Streptomycin (HLS - 300 µg) disk and High level Gentamicin (HLG - 120 µg) disk and Agar dilution method as per CLSI guidelines6 . As per CLSI guidelines, for detection of High Level Gentamicin Resistance by Agar dilution method, Gentamicin concentration was taken as 500 µg/ml. 10 µL of 0.5 Mc Farland suspension of test strain was spotted onto BHI (Brain Heart Infusion) agar surface containing Genatamicin 500 µg/ml6 . High Level Gentamicin Resistance (HLGR) was also confirmed by putting the Himedia Ezy MIC strip. A lawn culture of the test strain (turbidity adjusted to 0.5 Mc Farland) was done on Mueller Hinton (MH) agar plate and Himedia Ezy MIC strip was put and MIC (Minimum Inhibitory Concentration) was detected after incubation at 370C for 18 hours. MIC range of the strip was from 0.064-1024 µg/ml. E.faecalis ATCC 29212 was used as quality control for all the tests.
RESULTS
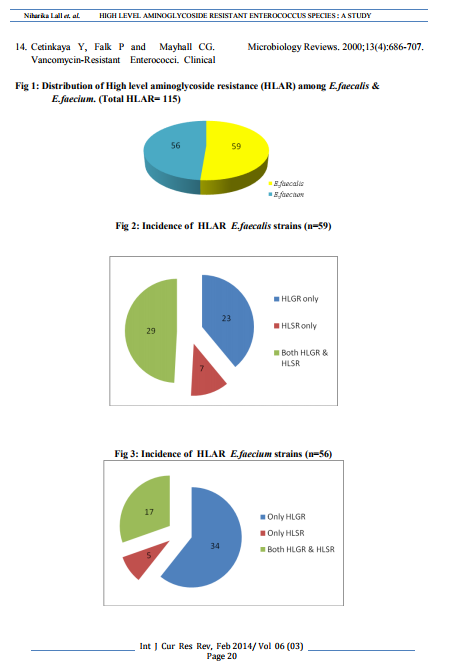
Out of 190 Enterococcus strains isolated, High level aminoglycoside resistance (HLAR) was detected in 115(60.5%) strains. HLAR strains were detected by disc diffusion test, agar dilution method and HLGR by Gentamicin Ezy MIC strip (photo1,2 and 3). Amongst 115 HLAR strains, 59 (51.4%) strains were E.faecalis and 56 (48.6%) were E.faecium (fig1). Only High level Gentamicin resistance (HLGR), only High level Streptomycin resistance (HLSR) and both HLGR and HLSR producing E.faecalis strains were detected as 23(38.9%), 7 (11.9%) and 29 (49.2%) respectively out of total 59 HLAR positive E.faecalis strains (fig 2). Similarly, out of total 56 HLAR producing E.faecium strains, 34(60.7%) produced only High level Gentamicin resistance (HLGR), 5(8.9%) produced only High level Streptomycin resistance (HLSR) and 17 (30.4%) produced both High level Gentamicin resistance (HLGR) and High level Streptomycin resistance (HLSR) (fig 3). Out of 93 HLAR Enterococcus strains isolated from urine samples, 49 (52.7%) were E.faecalis and 44 (47.3%) were E.faecium. 11(68.8%) HLAR strains were isolated from total 16 pus and wound swab samples. Out of 11 HLAR strains, E. faecalis and E. faecium were 7 and 4 strains respectively. Out of total 25 blood samples, 9(36%) strains produced HLAR and E. faecium strains were shown to exhibit more HLAR than E. faecalis. Amongst 9 HLAR Enterococcus strains, only 3(33.3%) strains were E. faecalis whereas 6(66.7%) were E. faecium. Only 2 HLAR strains were detected from other specimens, which were identified to be E. faecium (fig 4). Maximum 36 (31.3%) HLAR strains were isolated from Surgery ward followed by Pediatrics (22), Medicine (16) and Gynaecology ward (10) (fig 5). Out of 36 HLAR strains isolated from Surgery ward, 23 (63.9%) were E. faecalis and rest 13 were E. faecium. Only 7 (6.1%) HLAR strains were isolated from different ICUs (MICU-3, NICU-3 and PICU-1). No HLAR strain was isolated from OT ICU . Out of 115 HLAR strains, 90 Enterococcus strains were MDR (Multi-drug Resistant). MDR Enterococcus strains were detected on the basis of resistance (acquired) to Erythromycin, Tetracyline and High level aminoglycosides. All (100%) HLAR Enteococcus strains were sensitive to Vancomycin and Linezolid. Out of 115 HLAR strains, only 7 (6.1%) and 23 (20%) were sensitive to Quinupristin-dalfopristin and Ampicillin respectively. Nitrofurantoin sensitivity was tested for 93 HLAR Enterococcus strains isolated from urine sample only and 85(91.4%) HLAR strains were sensitive to Nitrofurantoin.
DISCUSSION
A common regime for treatment of serious enterococcal infections is the combination of cellwall inhibitors, such as penicillin, ampicillin or vancomycin; with aminoglycosides, such as streptomycin or gentamicin. The addition of cellwall inhibitor agent helps in the penetration of the aminoglycoside into the bacterial cytoplasm, making the intrinsically resistant organism aminoglycoside sensitive. The presence of HLAR in Enterococci makes the synergism of cell-wall inhibitor and aminoglycoside ineffective7 . In our study, 60.5% HLAR producing Enterococcus strains were isolated. Our study correlated well with Deshpande et al who had reported 58.8% HLAR producing Enterococcus strains8 . Mendiratta et al in 2008 reported 46% HLAR producing Enterococcus strains9 , whereas Vinod kumar et al in 2008 reported 65.6 % HLAR producing Enterococcus strains10. The study conducted by Mendiratta et al also reported that 69.6 % of their HLAR strains were E.faecalis and 30.4 % were E. faecium9 , whereas in the present study 51.4 % strains were E. faecalis and 48.6 % were E. faecium. As per χ 2 test, when only HLGR producing E. faecalis (23) and only HLGR producing E. faecium (34) are considered, we conclude, that isolation of only HLGR producing E.faecium is significantly higher than only HLGR producing E. faecalis11. In our study, out of 115 HLAR strains, 90 (78.3%) were MDR which is quite higher than reported by Deshpande et al (57%) in 20088 . MDR is defined as acquired nonsusceptibility to at least one agent in three or more antimicrobial categories12. Nitrofurantoin sensitivity was tested for 93 HLAR Enterococcus strains and it showed good activity (total 91.4% sensitivity) against both E.faecalis and E.faecium. Probably, the resistance could not be acquired by Enterococcal strains as Nitrofurantoin is not used commonly nowadays. HLAR (High level aminoglycoside resistance) i.e., Streptomycin MICs >2000 µg/ml or Gentamicin MICs > 500 µg/ml is an acquired resistance2 . HLAR occurs due to the presence of AME ( Aminoglycoside modifying enzymes). The most frequently encountered enzyme include dual function 2?phosphotransferase and 6?acetyl transferase conferring HLR to all available Aminoglycoside (Kanamycin, Gentamicin, Amikacin, Netilmicin, Tobramycin) except streptomycin13. Hence, gentamicin resistance is a good predictor of resistance to other aminoglycosides except streptomycin14 . 6?adenyl transferase is another AME which is responsible for HLR to streptomycin but does not inactivate other useful aminoglycosides13 .
CONCLUSION
We hereby conclude, that as High level Aminoglycoside resistant (HLAR) Enterococcus species can predict the loss of synergy between cell-wall active antimicrobial agents and Aminoglycosides, all Enterococcus strains must be screened routinely for HLAR by all Clinical Microbiology Laboratories to get an effective therapeutic outcome. Administration of Penicillin –Aminoglycoside combination without knowing HLAR status of the causative Enterococcus species can be fatal in life threatening Enterococcal infections.
ACKNOWLEDGEMENT
Authors are thankful to DMIMS (DU) for the help rendered, while conducting the study. Authors also acknowledge the immense help received from the scholars whose articles are cited and included in references of this manuscript. The authors are also grateful to authors/editors/publishers of all those articles, journals and books from where the literature for this article has been reviewed and discussed.
References:
REFERENCES
1. Tendulkar PM, Baghdayan AS, Shankar N. Pathogenic Enterococci: New developments in the 21st Century. Cell Mol Life Sci. 2003;60: 2622-2636.
2. Washington Winn, Jr., Stephen Allen, William Janda et al.Koneman’s Color Atlas and Textbook of Diagnostic Microbiology. 6th ed. Philadelphia: Lippincott Williams And Wilkins; 2006. Chapter 13, Gram –Positive Cocci Part II: Streptococci, Enterococci, and the “Streptococcus-Like” Bacteria; p.672-764.
3. Herman, D. J., and Gerding DN. Screening and treatment of infections caused by resistant enterococci. Antimicrob. Agents Chemother. 1991; 35: 215–219.
4. Papaparaskevas J, Vatapoulos A, Tassios PT, Avlami A, Legakis NJ and Kalapothaki V. Diversity among high-level aminoglycosideresistant enterococci. Journal of Antimicrobial Chemotherapy 2000;45: 277-283.
5. Facklam RR, Collins MD. Identification of Enterococcus species isolated from human infections by conventional test scheme. J Clin Microbiol. 1989;24: 731-734.
6. Clinical and Laboratory Standards Institute (2012) Performance standards for antimicrobial susceptibility testing: 22th informational supplement. CLSI document M100-S22. Clinical Laboratory Standards Institute, 940 West Valley Road, Suite 1400, Wayne, Pennsylvania 19087-1898 USA.
7. Adhikari L. High-level Aminoglycoside Resistance and Reduced Susceptibility to Vancomycin in Nosocomial Enterococci. J Glob Infect Dis. 2010; 2(3): 231–235.
8. Deshpande VR, Karmarkar MG, Mehta PR. Prevalence of multidrug-resistant enterococci in a tertiary care hospital in Mumbai, India. J Infect Dev Ctries. 2013;7 (2): 155-158.
9. Mendiratta DK, Kaur H, Deotale V, Thamke DC, Narang R, Narang P. Status of High level aminoglycoside resistant Enterococcus faecium and Enterococcus faecalis in a rural hospital of Central India. Indian Journal of Medical Microbiology. 2008; 26 (4): 369-71.
10. Vinodkumar C. S, Srinivasa H., Basavarajappa K. G., Geethalakshmi S., Bandekar N. Isolation of bacteriophages to multi-drug resistant Enterococci obtained from diabetic foot: A novel antimicrobial agent waiting in the shelf? Indian Journal of Pathology and Microbiology. 2011; 54 ( 1 ):90-95.
11. Park K. Park’s Textbook of Preventive and Social Medicine. 22nd ed. Jabalpur: Banarsidas Bhanot; 2013. Chapter 19, Health information and basic medical statistics: p.782-796.
12. Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, Harbarth S, Hindler JF, Kahlmeter G, Olsson-Liljequist B, Paterson DL, Rice LB, Stelling J, Struelens MJ, Vatopoulos A, Weber JT, Monnet DL. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012; 18(3):268-81.
13. Marothi YA, Agnihotri H, Dubey D. Enterococcal resistance:an overview. Indian Journal of Medical Microbiology. 2005; 23(4):214-219.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License