IJCRR - 6(24), December, 2014
Pages: 38-42
Date of Publication: 20-Dec-2014
Print Article
Download XML Download PDF
16S rRNA BASED T-RFLP ANALYSIS OF METHANOGENS IN BIOGAS PLANT WITH P. HYSTEROPHORUS L. AS SUBSTRATE
Author: Ramya R., Shree M. P.
Category: Healthcare
Abstract:Background: Methanogenic microbiome plays an important role in contributing to global renewable energy resource. Globally, biogas production totally estimates to 1/4th of total consumption of fossil fuel. The bioconversion of organic material by anaerobic digestion process to a valuable energy source has a promising outlook. Though there is lot of research contribution to the biogas technology, the knowledge about the microbiome involved in the bioconversion of plant biomass is limited. Objective: In this study, batch fermenter with P. hysterophorus as mono-substrate and along with co-additives of bovine animal faeces at different ratios is presented in the context of molecular data on the microbial composition. Material and methods: A slightly modified protocol of the kit RKN15 to extract the metagenomic DNA from the microbial community participating in the bioconversion of the mono-substrate P.hysterophorus, P. hysterophorus with cow dung and P. hysterophorus nwith goat dropping was performed. Thus extracted and purified DNA from the microbial consortium was subjected for 16S rRNA analysis by terminal restriction fragment length polymorphism (T-RFLP).
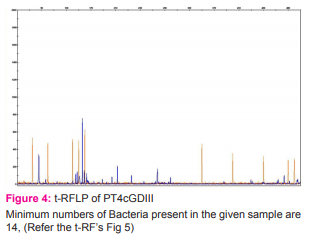
Result: The minimum estimate number of bacterial community present among the samples with high methane yield revealed the presence of 18 and 14 different types of microbes correspondingly.
Keywords: Bioconversion, Co-additive, Methanogenic, Microbiome, P. hysterophorus
Full Text:
INTRODUCTION
The diversity and functional analysis of the microbial community involved in the biogas technology is the principle aspect of upgrading the production efficiency in terms of methane yield. Hence a better understanding regarding the composition and abundance of the microorganisms involved in these systems becomes the need of the hour. Though there are reports of cultivable methanogens from diverse habitats (Garcia, 1990), cultivation of indigenous methanogens were unsuccessful (Williams and Crawford 1985; Goodwin and Zeikus, 1987). For the past few decades, researchers depend on culture independent methods of microbial community analysis using molecular techniques to isolate, identify and compare gene sequences (Atlas et. al., 1992; Madson, 1998; Schneegurt and Kulpa, 1998). Anaerobic digestion, an attractive option for the weed management at ambient environmental conditions offers technical and economical benefits. The potential limitations previously associated with weed biomass batch reactor operation have been largely overcome by optimizing various pretreatments and co-additive options. However, much scope still exists via application of the ambient microbial consortium which will allow in improvising the efficiency in terms of methane yield. In the present study, batch reactor fermenter operations were investigated by monitoring the effect of pretreatments and co-additives on reactor performance over hydraulic retention time of 35 days. DNA extracted from the whole microbial consortium was subjected to PCR amplification and 16S rRNA analysis by T-RFLP was performed.
MATERIALS AND METHODS
Batch Reactor Set-up
Biochemical Methane Potential (BMP) vials of 100ml were used as reactor bottles with P. Hysterophorus as substrate, which was pretreated (Ramya R and Shree M. P., 2014a). Inoculum substrate (I/S) ratio of 0.5%, 1.0%, 2.0% were done in triplicates. The inoculum was collected from a plant-litter based biogas plant, which was filtered with muslin cloth and stored at refrigeration temperature until use. Substrate along with co-additives (C/S ratio), cow dung and goat dropping at ratio of 1:0.25, 1:0.5 and 1:1 were performed in triplicates for a hydraulic retention time of 35 days in batch fermentation process at ambient temperature.
Biogas analysis
The composition of the biogas collected in the head space of the digester was analyzed using a Nano HP-1 Model BG 1000 Gas Chromatograph equipped with a Flame Ionization Detector (FID) and Thermal Conductivity Detector (TCD). Argon (Ar) was used as carrier gas with flow rate of 8ml/min. Oven temperature was set up to 60o C. Methanator temperature was set at 360o C. All the digesters were operated in triplicate. Biogas was sampled by directly inserting the intra venous syringe into the Anaerobic Digestion (AD) vial and volumetric composition of biogas was analyzed by using gas chromatography. Gas chromatograph (Nano HP-1) equipped with FID was used to analyze gas composition (Ramya R and Shree M. P., 2014b).
Extraction of metagenomic DNA from digested slurry
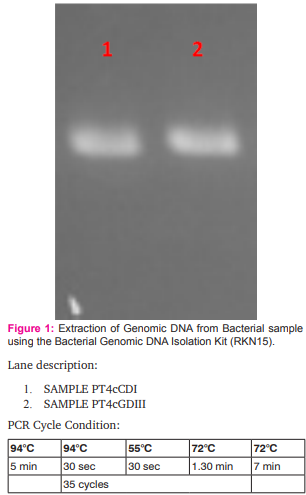
In this study, extraction of DNA of two samples PT4cCDI and PT4cGDIII were performed using the bacterial genomic DNA isolation kit RKN15 with slight modification. 1.5 ml of bacterial culture was taken in 2 ml vial after spinning the BMP vial at 70rpm for 10 mins. The supernatant thus obtained was spun at 10,000 rpm for 2 min. Supernatant was completely discarded. To the pellet 750 µl of 1X suspension buffer was added and subjected for vortexing. 5 µl of RNase was then added and incubated at 65°C for 10 min with intermittent mixing. To this, 1 ml of lysis Buffer was added and mixed by inverting the tube 10-12 times. The tubes were incubated at 65°C for 15 min and spun at 10,000 rpm for 5min. The clear lysate was loaded (unlysed Cells will settle as pellet leaving clear supernatant) onto spin column. The mini-spin column was placed into 2ml collection tube and 600µl of the lysate was loaded on the spin column each time, spun at 10,000 rpm for 1min at room temperature. The content of the collection tube was discarded and the spin column was placed back in the collection tube. 500 µl of 1X wash buffer was added to the column, spun at 10,000 rpm for 1 min at room temperature. The contents were discarded of the collection tube. The spin column was placed back in the same collection tube and the procedure was repeated. The empty column was spun at 10,000 rpm for 3 min at room temperature. 50 µl of warm elution buffer was added to the spin column placed in a fresh 1.5ml vial, at 65°C for 1 min and spun at 10,000 rpm for 1 min at room temperature. The elution was collected in the same vial. Thus obtained DNA concentration was determined by quantitative analysis on 1% (w/v) agarose gel with 0.5xTBE as electrode buffer as described elsewhere (Sun, et al., 2009).
T-RFLP analysis
The ~1.5 Kb bacterial 16s rRNA fragment was amplified from the metagenomic DNA extracts by using high–fidelity Polymerase Chain Reaction (PCR) polymerase as described by Sun, et. al., 2009, with the labelled primers marked with 6-carboxyfluorescein (FAM). The PCR products were subjected to restriction digestion with a 4-base cutter (HpaII). The fluorescent labelled t-RFs (terminal restriction fragments) were size separated on an ABI 3130 automated sequencer (Applied Biosystems) using an internal size standard (LIZ-500). t-RFLP (terminal restriction fragment length polymorphism) electropherograms were analyzed with GeneScan 3.7 software (Applied Biosystems) at chromous biotech Pvt Ltd, Bangalore (Wayan Suardana, 2014). The numbers of peaks obtained in t-RFs represents the minimum number of bacteria present in the sample.
RESULT
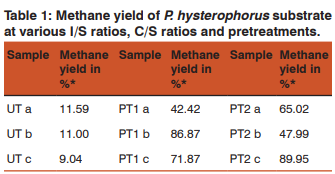
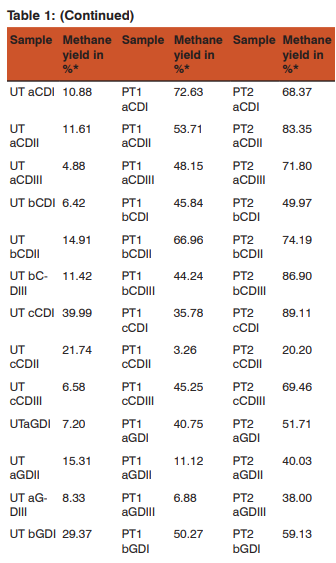
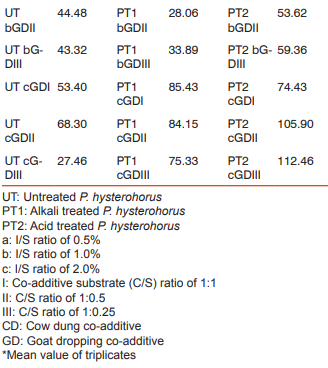
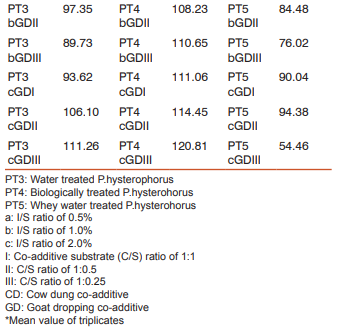
The biogas compositions of the samples subjected for AD are presented in Table 1 and 2. It is very evident from the GC result that the PT4 at the I/S ratio of 2%, C/S ratio of 1:1 with CD and C/S ratio of 1:0.25 with GD showed the maximum methane yield of 149.24% and 120.81%. Thus these two samples where further subjected for extraction of metagenomic DNA (Fig 1 and 2) and analysis by T-RFLP (Fig 3 and 4).
DISCUSSION
In our study, the DNA extracted from the BMP vials using slightly modified protocol of RKN15 kit gave us a good quality DNA which was amplified by PCR. T-RFLP is a technique widely used for study of microbial diversity in environmental samples (Salvador Embarcadero-Jimé- nex, et al., 2014). With the metagenomic DNAs obtained in this study, 35-500 T-RFs were obtained from the 16S rRNA amplicons digested by HpaII restriction endonuclease. Fig 3 and 4 indicated that the bacterial communities in the anaerobically digested P.hysterophorus present in the 100ml BMP vials were very diverse and composed of many species.
CONCLUSION
The effect of pretreatment and co-additive on the biomethanation efficiency of P.hysterophorus was studied by performing batch anaerobic digestion process experiment using BMP vials. The experimental results in this study demonstrated that the suitable pretreatment for biomethanation of P.hysterophorus for a HRT time of 35 days is PT4 with the maximum methane yield of 149.24% with CD as co-additive and 120.81% with GD as co-additive. The most important finding of this research is the microbial consortium involved in the biomethanation process. Therefore, the evaluation process with respect to the microbial consortium should be considered carefully. In the present study, extraction of high quality metagenomic DNA from anaerobically digested BMP vials was performed. Thus obtained DNA samples were subjected for PCR amplification and T-RFLP analysis, which showed the presence of diverse bacterial communities. Studies related to the diversity are still under progress.

6-carboxyfluorescein (FAM)-labeled primers gives blue color in Genescan analysis. (Blue colored peaks which corresponds to tRF’s are sized by internal size standard) Orange peaks are internal size standard (LIZ 500, Chromous)
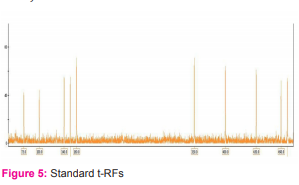
The GeneScan™ 500 LIZ® Size Standard is a fifth dyelabelled size standard for the reproducible sizing of fragment analysis data. This size standard is used for fragments between 35 and 500 bp. The standard contains 16 LIZ® dye-labelled, single-stranded DNA fragments. Since the standard is labeled with the fifth dye, can genotype a greater number of markers in a given lane, compared to the four-dye system.
Size Fragments in the 35-500 Nucleotides Range:
GeneScan™ 500 LIZ® Size Standard is designed for sizing DNA fragments in the 35-500 nucleotides range and provides 16 single-stranded labeled fragment of: 35, 50, 75, 100, 139, 150, 160, 200, 250, 300, 340, 350, 400, 450, 490 and 500 nucleotides. The sizing curve generated from these fragments make the GeneScan™ 500 LIZ® Size Standard ideal for a variety of fragment analysis applications such as Microsatellites, Fragment Length Polymorphisms and Relative Fluorescent Quantitation. Each of the DNA fragments is labelled with the LIZ® fluorophore which results in a single peak when run under denaturing conditions. With the 5th dye LIZ® your marker fragments can be labelled with the dyes FAM™, VIC™, NED™ or PET®.

ACKNOWLEDGEMENT
Authors acknowledge the immense help received from the scholars whose articles are cited and included in references of this manuscript. The authors are also grateful to authors/editors/publishers of all those articls, journals and books from where the literature for this article has been reviewed and discussed.
Source of Funding: No sources of funding
Conflicts of Interest: None
References:
1. Atlas R. M., Sayler G., Burlage R. S., and Bej A. K. Molecular approaches for environmental monitoring of microorganisms. BioTechniques, 1992;12:706-717.
2. Madsen E. L. Epistemology of environmental microbiology. Environ. Sci. Technol. 1998;32:429-439.
3. Garcia, J.L. Taxonomy and ecology of methanogens. FEMS Microbiol. Lett. 1990;87:297-308.
4. Goodwin S. and Zeikus J. G. Ecophysiological adaptations of anaerobic bacteria to low pH: analysis of anaerobic digestion in acidic bogs sediments. Appl. Environ. Microbiol. 1987;53:57-64.
5. Ramya. R, Shree. M. P. Comparative efficiency of pretreatment methods on Parthenium hysterophorus L., as a potential feed stock, “International Journal of Scientific and Research Publications, 2014.Vol. 4, Issue 9, September Edition”. pp 1-3 ISSN 2250-3153.
6. Ramya R, Shree M. P. Impact of C/N ratio of pretreated P. hysterophorus L., on Biomethanation, “Internation Journal of Scientific Research, 2014. Vol:III, Issue:XI, November Edition”, Journal DOI:10.15373/22778179.
7. Salvador Embarcadero-Jiménez, Feng Long Yang, Raquel Freye-Hernàndez, Yanelly Trujillo-Cabrera, Flor N. Rivera Orduña, Hong Li Yuan and En Tao Wang. An improved protocol for extraction of metagenomic DNA from high humus, alkaline and saline soil of Chinampa for T-RFLP fingerprinting analysis. British Microbiology Research Journal. 2014;4(7):821-830.
8. Schneegurt M. A. and Kulpa C. F. Review: The application of molecular techniques in environmental biotechnology for monitoring microbial systems. 1998;27:73-79.
9. Sun Y.M., Zhang N.N., Wang E.T., Yuan H.L., Yang J.S., Chen W.X. Influence of intercropping and intercropping plus rhizobial inoculation on microbial activity and community composition in rhizosphere of alfalfa (Medicago sativa L.) and Siberian wild rye (Elymus sibiricus L.) FEMS Microbiol. Ecol. 2009;70:62-70.
10. Wayan Suardana I. Analysis of Nucleotide Sequences of the 16S rRNA Gene of Novel Escherichia coli Strains Isolated from Feces of Human and Bali Cattle. Jornal of Nucleic acids, 2014. Vol (2014), Article ID 475745.
11. Williams R. T. and Crawford R. L. Methanogenic bacteria including an acid-tolerant strain, from peatlands. Appl. Environ. Microbiol. 1985;50: 1542-1544.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License