IJCRR - 14(1), January, 2022
Pages: 64-70
Date of Publication: 03-Jan-2022
Print Article
Download XML Download PDF
Oral Cancer Detection using Machine Learning and Deep Learning Techniques
Author: Nanditha B R, Geetha Kiran A Sanathkumar M P
Category: Healthcare
Abstract:Introduction: Oral cancer is one of the most dangerous cancers which occurs in the oral cavity. Overuse of tobacco and smoking cigarettes are the primary risk factors for developing oral cancer. Oral cancer diagnosis at an early stage can save the lives of many people with proper treatment. Objective: The proposed work aims at early detection of potentially malignant oral lesions by the development of an automated disease diagnosis system. Building a large dataset of well-annotated oral lesions is a primary key component. A novel strategy to build automatic oral cancerous image classification software is provided in this paper. Methods: In the present work, machine learning models and deep neural networks are used to build an automated diagnosis system. By using the initial data which was gathered in this study, Naive Bayes, KNN, SVM, ANN, and CNN classification models are constructed for the automated detection and classification of oral malignancies. A new CNN network is designed which consists of 43 deep layers, whose network structure is inspired by the standard VGG-16 network. Results: Performance analysis of different machine learning models and deep learning models has been provided. Results demonstrate that the deep learning model has the potential to tackle this challenging task of early detection of oral cancerous lesions. Conclusion: It is observed from experiments that different classifiers can perform well in identifying oral cancerous lesions. Particularly, the deep learning CNN model shows high accuracy in differentiating normal and cancerous images.
Keywords: Deep learning, Lesions, Machine learning, Texture features, Oral cancer, Convolution Neural Network, Benign, Malignant
Full Text:
INTRODUCTION
According to World Health Organization, nearly six lakh new cases of oral cancer and more than three lakh deaths are reported every year.1 Oral cancer include the main sub-sites of the lip, oral cavity, nasopharynx, and pharynx and have a particularly high burden in South Central Asia due to risk factor exposures.2 A comprehensive approach is needed for oral cancer to include health education and literacy, risk factor reduction and early diagnosis.
Computer vision can greatly assist in the diagnosis of oral cancer compared to a naked human eye examination, which turns out to be more complicated. Diagnosis and classification of oral cancer can be performed by traditional machine learning and deep learning techniques 3. In traditional machine learning techniques, a domain expert needs to identify the applied features and make them more clearly visible to the learning algorithm to work, to reduce the complexity. Literature reveals the use of conventional classification techniques like support vector machines (SVM), naïve Bayes, and k-nearest neighbor (KNN) classifiers for the classification of oral lesions into normal and abnormal lesions.4
SVM is a supervised machine learning algorithm that can be employed for classification.5 A method called the kernel trick is used to modify the data and then an optimal boundary is chosen. Naive Bayes is mainly employed in the classification problem and uses a similar technique which is probabilistic based on different features of different classes.6 K-nearest neighbor is used in predicting the class label of unknown data by taking the majority of the class labels of K-nearest data instances and the distance between the instances is measured by various distance measures 7.
Deep learning is a subfield of machine learning which works on artificial neural networks, which are algorithms inspired by the structure and function of the brain. ANN (Artificial Neural Network) can be trained on numerous images of both benign and malignant lesions. By learning the non-linear interactions, the model can tell itself if the image is malignant or benign. So, in deep learning, there is no need for domain expertise for feature extraction. In the present work, the classification of oral images into benign or malignant images is performed by a deep learning model using CNNs (Convolutional Neural Networks), which is a type of ANN.
CNN's are composed of multiple layers of artificial neurons. The behavior of each neuron is defined by its weights. Every CNN network consists of 4 layers: input layer, convolution layer, pooling layer, and fully connected (FC) layer. The convolutional layer extracts feature maps from the input image using filters and the pooling layer replaces the output of the network at certain locations by deriving a summary statistic of the nearby outputs. This leads to reduction of spatial size and thus reduces computational complexity. Neurons in fully connected layers have full connectivity with all neurons in the preceding and succeeding layers. The FC layer helps to map the representation between the input and the output. To classify the output, generally, a softmax layer is used in image classification. CNN's are trained using labeled datasets given with the respective classes. CNN's learn the relationship between class labels. For the input image shown in Figure 1, the feature maps extracted by the convolutional layer are shown in Figure 2.
LITERATURE REVIEW
Various researchers have tried different machine and deep learning techniques for the classification of images into normal and abnormal images. Licheng Jiao et al.8 present a survey on new-generation deep learning techniques which can be used for image processing tasks. Three series of deep learning models namely, CNN series, GAN series, and ELM series networks and their roles in image processing tasks have been described. These are used extensively in image processing nowadays, where these techniques have different depth and types of networks which makes image processing tasks easier now. Daisuke Komura et al. 9 have applied machine learning techniques like SVM, random forest, CNN, k-means, auto encoder, and principal component analysis for histopathological image analysis. Before applying machine learning methods, feature extraction and classification between cancer and non-cancer patch are performed.
A survey on feature extraction methods that extract meaningful features from the raw images has been presented by Anne Humeau et al.10 Seven classes of texture feature extraction methods, their advantages, disadvantages, and applications are reviewed. A large number of texture datasets that are used by the authors to test and compare the feature extraction algorithms have been described. For very high-resolution remote sensing images, the authors have proposed histogram-based attribute profiles that allow the modeling of texture information from attribute profiles.
A discussion on supervised, unsupervised, and semi-supervised feature selection techniques which reduce computation time, increase accuracy and help in removing redundant and irrelevant data has been presented by Jie Cai et al.11 Their use in many fields like image retrieval, text mining, fault diagnosis, and other areas has been reviewed by the authors.
Skin segmentation using Yellow-Chrominance blue-Chrominance red (YCbCr) and Red-Green-Blue (RGB) color models is presented by Shruti et al.12 The authors have developed a computationally efficient and accurate approach for skin cancer detection which may be used in real-time. Results indicate that the YCbCr model is better than the RGB model in segmenting and classifying skin lesions.
A deep learning model has been developed by the authors for classifying oral malignant lesions by the use of CNN technique.13
DATA COLLECTION
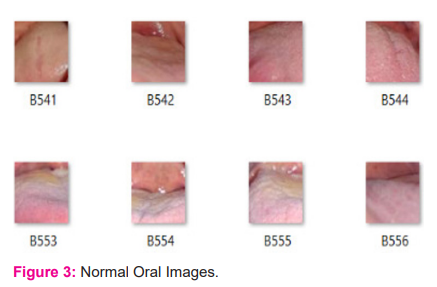
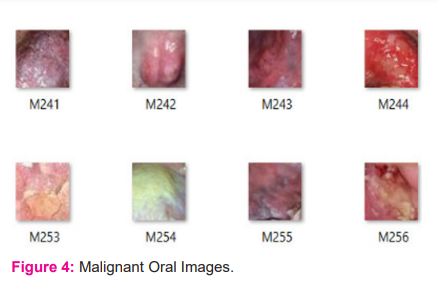
A total of 630 oral images have been used in the present work for oral image classification. Few of these images were downloaded from the internet and few others were collected from different hospitals by consulting oral specialists. From these images, 1200 lesion regions were cropped and as a result, we obtained separate images of lesion regions. Out of those 1200 lesion images, 600 are malignant images and 600 are normal images. Figure 3 and Figure 4 depict different patches of few normal and malignant images from the dataset. The benign patches are labeled as B001, B002, and so on. The malignant patches are labeled as M001, M002, and so on. A feature vector was created by extracting useful features from these lesion images and then the newly obtained dataset is used in machine learning models for testing purposes. For the deep learning model, the lesion images are augmented to generate 9600 images. Then these images are used for training the network.
MATERIALS AND METHODS
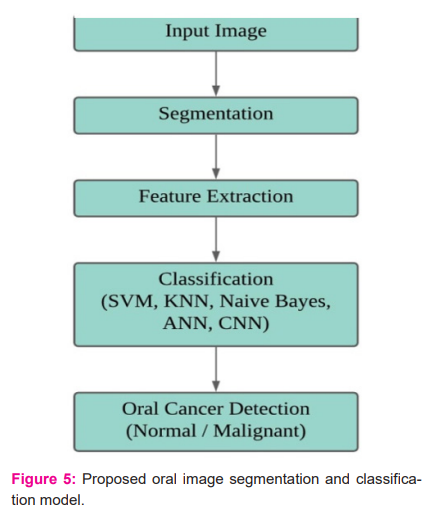
The details of the proposed work for oral malignancy detection have been elucidated in Figure 5. The input to the disease diagnosis system is an RGB oral image; this input image is then subjected to a segmentation process in order to select the lesion region. After segmentation, features are extracted from the lesion region; these features are used to classify the image using SVM, KNN, Naive Bayes, and ANN models. CNN takes a segmented lesion image as an input to the network and classifies the image as normal or malignant. The details of this process are elaborated in the following sections.
Segmentation
In this process, the lesion region is extracted from an input image. Firstly, the input RGB image is converted into YCbCr color space and then a mask of lesion region is created based on blue difference chroma (Cb) and red difference chroma (Cr) intensity values. Cancerous oral images have two types of lesion patches-white and red lesion patches.
If the mean Cr value of the input image is less than a predefined threshold value, then that image will have white patches; a mask of white lesion patches is created using Cb intensity, if a region contains a mean Cb value more than the mean Cb value of the whole image then that region is considered as a white lesion patch. If the mean Cr value of the input image is greater than the predefined threshold value, then the input image will have red patches; a mask of red lesion patches is created using Cr intensity, if a region contains a mean Cr value more than the mean Cr value of the whole image then that region is considered as a red lesion patch.
After creating the lesion mask, a region-based active contour segmentation is performed, which selects the lesion regions in the input image based on the mask, and the lesion which has the wider area is considered as the final output lesion and then that lesion is extracted as a separate image. Figure 6 depicts the details of oral lesion segmentation.
Feature Extraction
A feature vector of 44 features is created by extracting Grey-level co-occurrence matrix (GLCM), Grey-level run-length matrix (GLRLM), Fractal features, Gabor features, and Color features from lesion images. The extracted features are Energy, Homogeneity, Contrast, Correlation, Short-run emphasis, Long-run emphasis, Gray-level non-uniformity, Run-length non-uniformity, Run percentage, Low gray-level run emphasis, High gray-level run emphasis, Short-run low gray-level emphasis, Short-run high gray-level emphasis, Long-run low gray-level emphasis, Long-run high gray-level emphasis, Fractal dimension, Fractal lacunarity, Standard deviation, Gabor mean squared energy, Gabor mean amplitude, Mean and Standard deviations of RGB, Hue-Saturation-Value (HSV) and YCbCr color components.
Feature Selection
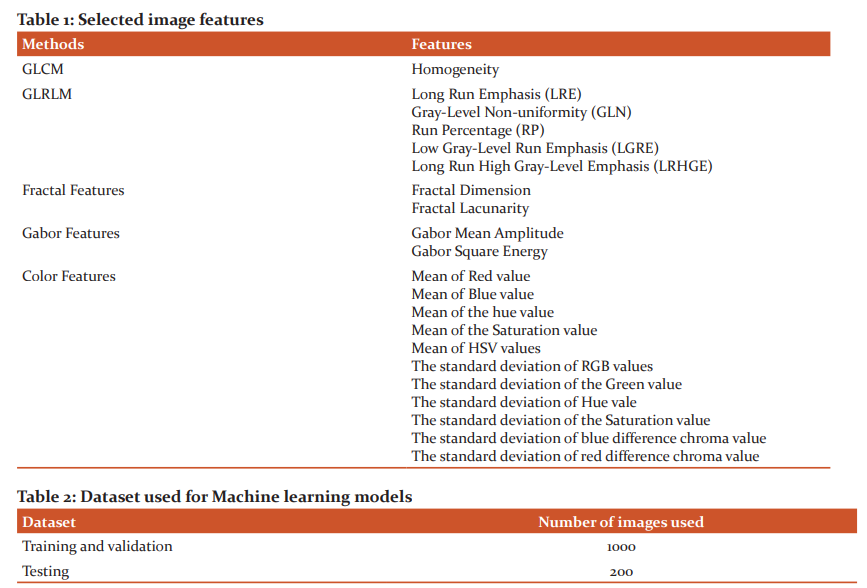
The irrelevant and redundant features from extracted features have been removed and only the most relevant features are selected by using the statistical feature selection methods: Minimum Redundancy Maximum Relevance (MRMR) and Box plot methods. Based on the rank assigned by these feature selection methods, the top 19 features are selected. The selected features are listed in Table 1.
Classification
The dataset of 19 selected features is used for training machine learning classification models and the ANN model. Segmented lesion images are used for training the CNN model. The classification Techniques employed in this paper are described below:
Support Vector Machine (SVM)
SVM is a supervised machine learning algorithm that can be employed for the classification process. Medium Gaussian SVM model is used for classification which utilizes a method called the kernel trick to modify the data and then based on these changes it identifies an optimal boundary among the possible output.
K-Nearest Neighbour (KNN)
A modified version of KNN i.e., weighted KNN is used to classify the image based on selected features from the lesion region. The performance of this method is completely dependent on the training set and the choice of hyperparameter K. For the current work, a value of 10 is chosen for K.
Naïve Bayes
A Naive Bayes classifier is a simple probabilistic classifier that is based on the Bayes theorem. For the present classification, a Bayesian classifier with different kernel densities is made use of.
Artificial Neural Network (ANN)
A Feed-Forward Artificial Neural Network is used to classify images based on selected features from the lesion image. A dataset of 19 features extracted from lesion images is used to train the network. The network consists of 20 hidden layers and 1 output layer. The ANN network architecture is shown in Figure 7.
Convolutional Neural Network (CNN)
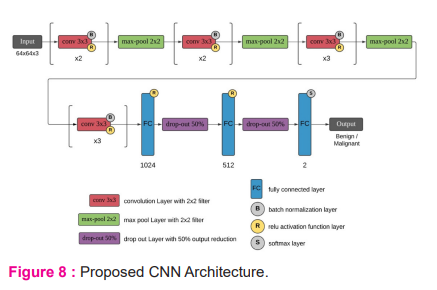
A new convolutional neural network is created for oral disease detection, which has 43 layers. The architecture of the CNN network is shown in Figure 8. This network includes 10 convolutional layers, Batch normalization layer is used to normalize convolutional layer output and the ReLU layer is used activation function. The max-pool layer is used for pooling. 3 fully connected layers are used with 1024, 512, and 2 nodes in respective layers. Output is predicted using softmax layer. This CNN network takes 64x64 RBG images as input and classifies them as benign or malignant images.
RESULTS
Dataset
A dataset that consists of 19 feature values of 1200 lesion images is used to train using SVM, KNN, Naive Bayes, and ANN models. Training and testing dataset details for the used models are given in Table 2. Segmented lesion images are used for training and testing the CNN model and their dataset details are given in Table 3.
Training and Testing results
The training accuracies of the classification models are given in Table 4. The testing performance measures of the classification models are given in table 5.
DISCUSSION
From the obtained results, among machine learning classification models i.e. Naive Bayes, KNN and SVM, the SVM model performs well when compared to the other two models. CNN model performs better when compared to ANN. Though ANN’s performance is nearly equal to CNN, CNN still stands out to be best because of its ability to learn better features by itself by looking at the overall performances of each classification model, CNN outperforms all other models with 99.3% training accuracy and 97.51% testing accuracy. Also ANN and SVM model performs almost equally when considered to the performance of CNN.
CONCLUSION
In this paper, a few machine learning and deep learning classification models for oral cancer detection have been discussed. The results of the classification models for automating the early detection of oral cancer have been demonstrated. Desktop software is built using Matlab to classify whether an image is malignant or normal by using any of the models which are presented in the paper and a report is generated accordingly.
The promising model results demonstrate the effectiveness of deep learning and suggest that it has the potential to tackle these challenging tasks.
FUTURE SCOPE
In future work, it will be possible to gather more images for enriching the dataset and to improve the accuracy of the models using different techniques of fine-tuning and augmentation. The main goal will be implementing a semantic segmentation for selecting lesion region from an input image to improvise accuracy results of the models.
Acknowledgment
The authors acknowledge the immense help received from the scholars whose articles are cited and included in references of this manuscript. The authors are also grateful to authors/editors/publishers of all those articles, journals, and books from where the literature for this article has been reviewed and discussed
Source of Funding
No financial support has been obtained for this work
Conflict of interest
Authors have no conflict of interest
Author’s Contribution
All authors have substantially contributed to the conception and design of the manuscript and interpreting the relevant literature. The authors have drafted the manuscript and revised it carefully for important intellectual content.





References:
-
https://www.cancer.org/cancer/oral-cavity-and-oropharyngeal-cancer/about/key-statistics.html.Key Statistics for Oral Cavity and Oropharyngeal Cancers. American Cancer Society.
-
https://www.cancer.org/cancer/oral-cavity-and-oropharyngeal-cancer/causes-risks-prevention/riskfactors.html. Risk Factors for Oral Cavity and Oropharyngeal Cancers. American Cancer Society.
-
Dargan S, Kumar M, Ayyagari M R. A Survey of Deep Learning and Its Applications: A New Paradigm to Machine Learning Arch. Computat Methods Eng 2020; 20: 1071–1092.
-
Prabhakaran R, Mohana J. Detection of Oral Cancer Using Machine Learning Classification Methods. Int J of Elect Eng and Tech 2020; 11(3): 384-393.
-
Shujun H, Nianguang C, Pedro P, Shavira N, Yang W, Wayne X. Applications of Support Vector Machine (SVM) Learning in Cancer Genomics. Cancer Genomics & Proteomics 2018; 15: 41-51.
-
Khanna D, Sharma A. Kernel-Based Naive Bayes Classifier for Medical Predictions. Adv in Int Sys and Compvol 695 Springer Singapore.
-
Lavanya L, Chandra J. Oral Cancer Analysis Using Machine Learning Techniques. ISSN 0974-3154, Volume 12, Number 5 2019: 596-601.
-
Licheng J, Jin Z. A Survey on the New Generation of Deep Learning in Image Processing.IEEE Access 2019; 7: 172231-172263.
-
Daisuke K, Shumpei I. Machine Learning Methods for Histopathological Image Analysis.Compt and Struct Biotec J, 2020; 16: 34-42.
-
Anne H-H. Texture Feature Extraction Methods: A Survey. Journal of IEEE Access, 2019;7:8975-8999.
-
Jie C, Jiawei L, Shulin W, Sheng Y. Feature selection in machine learning: A new perspective. Neurocomputing Journal Elsevier, 2018; 300: 70-79.
-
Shruti DP, Wayakule JM, Apurva DK. Skin Segmentation Using YCbCr and RGB Color Models. Int J of Adv Res Comp Sci and Soft Eng, 2014; 4(7).
-
Welikala RA. Automated Detection and Classification of Oral Lesions Using Deep Learning for Early Detection of Oral Cancer. IEEE Access, vol. 8, 132677-132693, 2020.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License