IJCRR - 13(19), October, 2021
Pages: 14-16
Date of Publication: 11-Oct-2021
Print Article
Download XML Download PDF
Knowledge Discovery in Protein Sequence Analysis Using Hierarchical Clustering Method
Author: Desai Farhana
Category: Healthcare
Abstract:Introduction: New Data Mining techniques play a very important role in the large growing biological databases. The clustering technique is an unsupervised method in data mining. Hierarchical Clustering techniques are useful to represent relationships between protein families. Objective: Bioinformatics urges the need of discovering knowledge in the vast area of molecular biology by using data mining as the core. Data Mining aims to discover hidden data from a large volume of data. Method: This paper discusses the hierarchical clustering technique of data mining on protein sequence datasets to identify genes that are consistent, easy to implement, finding and grouping the number of clusters pattern recognition. However, the valuable data is sometimes not useful but the knowledge hidden in that valuable data is meaningful. Result: The distance is used to determine how closely two organisms are related, whereas the dendrogram shows a graphical representation of the distance calculated between the clusters. Conclusion: The hierarchical clustering will help the biologist to judge which genes were clustered rightfully by viewing the tree structure. e the dendrogram. Therefore, the main aim is to unfold the knowledge in the vast field of bioinformatics by using information technologies as the key.
Keywords: Clustering, Phylogenetic tree, Sequence, Hierarchical clustering, Pattern, Dendrogram
Full Text:
INTRODUCTION
Bioinformatics and data mining provide exciting research challenges for computational science. Data Mining is the process of knowledge discovery of various patterns from a sea of data whereas Bioinformatics involves the storage, analysis, and construction of information from biological data in the form of sequences, pathways, and gene expression. New Data Mining techniques play a very important role in the large growing biological databases. Many times Data Mining is also known as Knowledge Discovery, which means searching a large volume of data to discover patterns and new trends that go beyond simple analysis.1
The Clustering technique is an unsupervised method in data mining.2 It helps to improve the accuracy of the data. A phylogenetic tree can be build-using Hierarchical clustering for known corresponding protein sequences or DNA sequences. A Phylogenetic tree is useful for biologists in solving both scientific and practical problems. The problems would be stated as an evolution of complex features of the species, prediction about fossils, the evolution of diversity, etc.
Hierarchical Clustering techniques are useful to represent relationships between protein families. There are two types of hierarchical clustering methods Agglomerative (bottom-up) and Divisive clustering (top-down). Using agglomerative clustering the very first single cluster is chosen and further in recursive fashion two or more relative clusters keep on merging. Divisive clustering starts with all data points merged in one cluster and further splits into most relative clusters recursively. Cluster analysis is a method for finding similar data objects present in the data.3This method divides the section into sets of clusters in such a way that two objects a picked, which resembles similar from the same cluster, whereas clusters, which belong to different clusters, are not similar.
METHODOLOGY
The process of data mining is part of many steps, which require repetition and refinement to achieve accuracy. Finding out new methods of Data Mining will act as a key factor in the vast mounting of biological data. It is a key point to state that the whole process of Data Mining covers a gathering of many techniques [Fig. 1], which involves finding the pattern from a large set of sequences, applying a hierarchical clustering algorithm and discovering knowledge through the result, which is a phylogenetic tree.
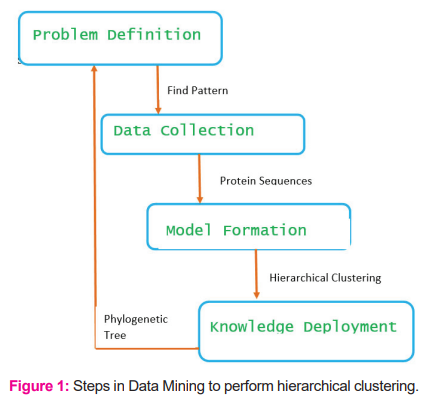
CONCLUSION
The phylogenetic tree is compared against the problem if it is solved with a single phylogenetic tree then there is no recursive procedure to be followed else the data is changed and the hierarchical clustering steps are executed to get the right phylogenetic tree.4 The data is collected from a distance matrix [Fig. 2] on which the hierarchical clustering method is applied.5
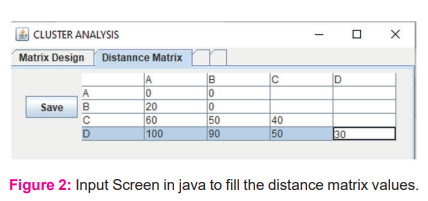
The output of Hierarchical clustering is a dendrogram that is built from a cluster hierarchy or, in a different way, a tree of clusters. Each cluster node arises from child clusters; sibling clusters extract the points roofed by their common parent. Such a method allows discovering data on various granularity levels. The process continues in this form until the requested number of clusters i.e. a constant k is achieved.
RESULTS
The closest clusters were displayed through the hierarchical clustering algorithm as follows:
The closest Clusters are:
C, DE:: 45
A, B:: 20
AB, CDE:: 72
D, E:: 30
The distance is used to determine how closely two organisms are related. The line that connects
all other lines is the representation of the common ancestor. Fig.3 describes the dendrogram generated from the distance matrix.
DISCUSSION
A dendrogram is a cluster tree, which is used to represent sequence data providing links to various groups. Each group is connected to a similar group and ends up forming a tree-like structure.
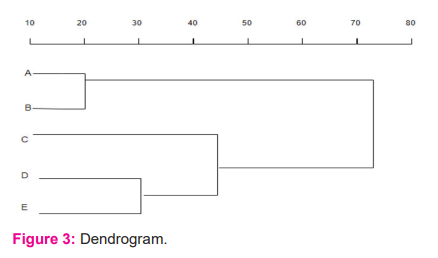
The above figure shows the results of a hierarchical cluster analysis of the sequence. The horizontal axis shows the distance between each cluster, in which four clusters were identified with an optimal number of 2–4nodes in each cluster. Cluster 1 is a combination of A and B, which are the closest cluster with a distance of 20. The next closest found is D and E with a distance of 30. The common factors for C and (D and E) are grouped in Cluster 3, whereas Cluster 4 is a combination of (A and B) and (C and (D and E)). This information, which is divided amongst the nodes, has a huge impact on the selection of the closest cluster.
The advantage of hierarchical clustering helps the biologist to judge which genes were clustered rightfully by viewing the tree structure. e the dendrogram. It also enables us to predict how the genes will change in the future. Phylogenetics provides a more accurate description of patterns based on sequence data.
Acknowledgement: I am thankful to Dr. Acharya for productive discussion in the field of bioinformatics and computer science. Sir, guided me to include the best of computer skills in the study of bioinformatics. I also thank my Institute colleagues for their healthy discussion that supported my research.
Conflict of interest: The author declares that there are no competing interests
Source of Funding: No source of funding
Authors’ Contribution: The author has proposed the approach and has also read and approved the final manuscript
References:
1. Vinothini B, Shobana D, Nithyakumari P. Application of Data mining in the Field of Bioinformatics. Int J Trend Res Dev. 2016. 3(1):21-3.
2. Ali Masood M, Khan MNA. Clustering techniques in bioinformatics. I.J. Modern education and computer science.Int J Med Curr Sci. 2015; 1:38-46.
3. Murugananthi C, Ramyachitra D. Performance evaluation of partition and hierarchical clustering algorithms for protein sequences. Int J Comp Intell Inform Int J Curr Infor. 2014;3(4):272-7.
4. Staton JL. Understanding phylogenies: constructing and interpreting phylogenetic trees. J South Car Acad Sci. 2015;13(1):24-9.
5. Desai F, Kamat RK. Calculation and visualization of the phylogeny of clusters using java API (application programming interface). J Eng Appl Sci. 2017;12(11):2827-30.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License