IJCRR - 13(17), September, 2021
Pages: 136-139
Date of Publication: 12-Sep-2021
Print Article
Download XML Download PDF
Efficacy of Modified Carbapenem Inactivation Method for Carbapenemase Detection and Comparative Evaluation with Polymerase Chain Reaction for the Identification of Carbapenemase-Producing Klebsiella pneumonia Isolates
Author: Bhat Asifa, Benazir Shazia, Fomda Bashir Ahmad
Category: Healthcare
Abstract:Introduction: Rapid and reliable detection of carbapenemase-producing microorganisms is an important element of antimicrobial stewardship. Various phenotypic methods for the detection of these organisms have their limitations. Aims: This study was conducted to assess the performance of the Modified Carbapenem Inactivation Method (mCIM) which is a recent specific phenotypic method described by CLSI 2018 guidelines for the detection of carbapenemase production. Methodology: In the study, mCIM was performed in 80 Gram-negative isolates that were resistant to meropenem by the Kirby�Bauer disc diffusion method. Separately, another 20 blaKPC( Klebsiella pneumoniae carbapenemase) polymerase chain reaction (PCR) positive and 15 blaKPC PCR negative Klebsiella pneumoniae isolates were tested by mCIM and their results were compared. Results: The positivity of mCIM was 92.7% and 80% in the case of Enterobacteriaceae and Pseudomonas spp. respectively. The test also showed 100% sensitivity for detecting carbapenemase production in PCR positive Klebsiella pneumoniae isolates harbouring the blaKPC gene. Conclusion: mCIM is a user-specific phenotypic test that is simple and easy to carry out for the detection of carbapenemase-producing microorganisms in microbiology laboratories.
Keywords: Carbapenem resistance, Carbapenemases, mCIM, Phenotypic method, Polymerase chain reaction
Full Text:
INTRODUCTION: Carbapenemases have become a frequent cause of drug resistance in Enterobacteriaceae, Pseudomonas aeruginosa and Acinetobacter baumannii during the last decade.1 Multiple carbapenemase genes have been described till now. The common determinants contributing to carbapenem resistance include blaKPC (Ambler class A), blaNDM(Ambler classB), and blaOXA-48-like(Ambler class D) genes. These enzymes are routinely isolated from Klebsiella pneumoniae and Escherichia coli along with other Gram-negative non-fermenters like Pseudomonas aeruginosa and Acinetobacter spp.2 Infections due to carbapenemase-producing Gram-negative bacteria can cause serious illness leading to a prolonged period of hospitalization and increased mortality ratio. Therefore, monitoring the development of resistance against carbapenems is of utmost importance.3
Various phenotypic, as well as genotypic methods, have been used to detect carbapenemases. Modified Hodge test was a useful phenotypic test previously included in the CLSI guidelines but it had the disadvantage of giving false-positive results mostly with Enterobacter spp. harbouring AmpC enzymes and alterations in porin channels. It also suffered false-negative results in New Delhi Metallo-beta-lactamase(NDM) producing isolates.4 Other phenotypic tests like the Carba NP test requires the use of specific and exclusive reagents, also the interpretation of the Carba NP test is subjective and it has shown poor sensitivity for the detection of OXA-48-type carbapenemases.5,6 Genotypic assays can principally detect only known targets and thus the performance of such test could be seriously affected by the occurrence of mutations within the targets.4
Modified carbapenem inactivation method (mCIM) is included as a screening test for carbapenemase-producing Enterobacteriaceae in CLSI 2018.7 Compared to Carba NP, which has a rapid turnaround time of 2 hours, mCIM is time-consuming, requiring an overnight incubation for the detection of carbapenemases. But it is relatively simple and has shown sensitivity and specificity of more than 99%.8
The emergence and spread of carbapenem-resistant bacteria is an issue of great clinical and public health concern.4 Thus, the study was carried out to evaluate the usefulness of mCIM for the identification of carbapenem-resistant isolates in our hospital because the accurate diagnosis of infections due to carbapenemase-producing Gram-negative bacteria is important for epidemiological purposes, infection-prevention measures and expediting appropriate therapy in such infected patients.
MATERIAL AND METHODS
This prospective cross-sectional study was conducted in the Department of Microbiology, Sher-i- Kashmir Institute of Medical Sciences (SKIMS) India, a tertiary care institute from October 2019 to March 2020.
Clinical Isolates: 80 Gram-negative isolates recovered from various clinical samples (blood, pus, urine, sputum and other body fluids) were included in the study. The isolates comprised of Escherichia coli, Klebsiella pneumoniae and Pseudomonas aeruginosa.
Antibiotic susceptibility was performed by Kirby-Bauer disc diffusion method according to CLSI guidelines and the isolates resistant to meropenem(10 µg) were selected.7 These isolates were subjected to a modified carbapenem inactivation method(mCIM). In addition, 20 blaKPC PCR positive and 15 blaKPC PCR negative Klebsiella pneumonia isolates were tested by mCIM separately.
mCIM testing: It was done according to CLSI 2018 guidelines.7 Using a sterile inoculating loop, 1µl of the test organism(Escherichia coli / Klebsiella pneumoniae) or 10µl of Pseudomonas aeruginosa was added into a tube containing 2 ml of tryptic soy broth(HIMEDIA); the bacterial suspension was vortexed for 10 to 15s. Aseptically, a 10µg meropenem disc (HIMEDIA) was added to this bacterial suspension and was incubated for 4 h ±15 min at 35°C ± 2°C in ambient air. Preceding the completion of the 4-hour carbapenem inactivation step, a 0.5 McFarland standard suspension of the mCIM indicator organism (Escherichia coli ATCC 25922, a carbapenem-susceptible strain) was prepared and inoculated on the Mueller-Hinton agar plate(HIMEDIA) using the method for standard disc diffusion susceptibility testing. The meropenem disc was then removed from the tryptic soy broth bacterial suspension using a 10µl inoculating loop which was dragged along the edge of the tube to remove excess liquid, and the disc was placed on the inoculated MHA plate. It was then incubated for 18 to 24 h at 35°C ± 2°C in ambient air.7
mCIM result interpretation: The test was interpreted according to CLSI 2018 guidelines.7 The zone of inhibition around each meropenem disc was measured. A zone diameter of 6 to 15 mm or presence of colonies within 16-18mm zone was considered as a positive result (i.e. carbapenemase production ), a zone diameter of more than or equal to 19 mm was considered a negative result (i.e. no carbapenemase production ) and a zone diameter of 16-18 mm was considered an indeterminate result.7 A small ring of a growth adjacent to the meropenem disc, representing carryover of the test organism from the Tryptic soy broth was ignored (Fig.1). For intermediate results, mCIM for the test isolate was repeated after checking for the purity of Escherichia coli ATCC 25922 indicator strain and the meropenem disc integrity by subjecting it to repeat disc diffusion test.

Polymerase chain reaction (PCR): Molecular identification of KPC-producing Klebsiella pneumoniae was done by bla KPC PCR using bacterial lysates prepared by removal of 200μl of overnight broth culture. The lysates were centrifuged at 12,000 × g for 2 min, and then re-suspended in 200μl of molecular-grade water followed by boiling at 95°C for 10 min, and discarding the cellular debris by centrifugation at 12,000 × g for 2 min at 4°C. 1μl of cell lysates were then subjected to PCR analysis using the following primers designed to identify all blaKPC genes (blaKPC-1 through blaKPC-7):
-
KPC forward (ATGTCACTGTATCGCCGTCT),
-
KPC reverse (TTTTCAGAGCCTTACTGCCC).
The reaction was set up in a PCR vial, after the addition of master mix, primers and the extracted DNA. 25μl of Master Mix contained 10X Taq buffer, 0.4mM dNTPs mix, 2mM Mgcl2, and 2U Proofreading Taq DNA polymerase.(Thermo scientific, USA). Lysates derived from blaKPC carrying Klebsiella pneumoniae strain 1705 and Escherichia coli ATCC 25922 were used as positive and negative controls respectively. The PCR was set up at the following conditions: 15 min at 95°C and 38 cycles of 1 min at 94°C, 1 min at 62°C, and 1 min at 72°C, which was followed by an extension step of 10 min at 72°C.9 The PCR products were analysed by electrophoresis on 2% agarose gel stained with ethidium bromide and visualized with UV light. The blaKPC gene gave a band at 893bp.
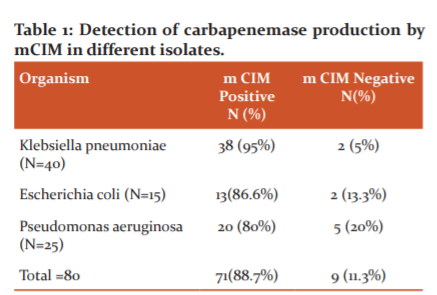
RESULTS: Out of the total 80 isolates tested for carbapenemase production by mCIM, 71 (88.7%) were carbapenemase producers. Initially, an indeterminate result was present in 12 isolates(8 Klebsiella pneumoniae and 4 Pseudomonas aeruginosa isolates). After repeating mCIM of these isolates 6 among 8 Klebsiella pneumoniae isolates were mCIM positive and 2 were mCIM negative. Among the 4 Pseudomonas aeruginosa isolates, 2 were positive and 2 were negative after repeating mCIM. The positivity of mCIM in different isolates is shown in Table-1.
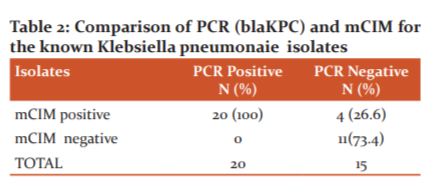
While comparing mCIM of the blaKPC Klebsiella pneumoniae isolates, it showed 100% sensitivity in the detection of carbapenemase production in PCR positive Klebsiella pneumoniae isolates harbouring the bla KPC gene. (Table 2)
DISCUSSION
Resistance to carbapenems is a worrisome international public health problem.2 Rapid detection of carbapenemase-producing Enterobacteriaceae is important both for taking effective precautions against the infection and for starting the appropriate treatment procedure.3 Polymerase chain reaction is considered the gold standard for carbapenemase gene identification, but it has some limitations. False-negative PCR results can occur due to a carbapenemase gene not tested in the PCR reaction, or mutations affecting the annealing of primers. False-positive PCR results can be due to the detection of inactive genes (with no carbapenemase expression). Such results can delay infection-control measures or, oppositely, initiate them when they are not required.10
Different phenotypic methods also have their shortcomings. The sensitivity and specificity of these tests vary depending on the bacterial species, enzyme type and expression level of the gene that codes for the enzyme or carbapenem resistance.3
In our study, we evaluated the performance of the mCIM (recommended by CLSI 2018) which is a simple test and has been seen to have better sensitivity and specificity than other phenotypic methods for the detection of carbapenemases.
The overall positivity of mCIM for Enterobacteriaceae and non-Enterobacteriaceae (Pseudomonas spp.) in our study was 88.7%. 11.3 % of our isolates that were resistant to meropenem by disc diffusion were mCIM negative which could be because of the various other
mechanisms of resistance in these isolates rather than carbapenemase production. It was observed that the detection of carbapenemases was high (92.7%) in the case of Enterobacteriaceae (Klebsiella pneumoniae and Escherichia. coli). The overall sensitivity of mCIM for Enterobacteriaceae is 93% to 100% in a study by Pierce VM et al. 4 In the case of Pseudomonas isolates, the positivity of mCIM in our study was 80%. In a study by Luiz F et al., the sensitivity of mCIM was 81% for Pseudomonas isolates.11 In another multicentric study by Simner PJ et al. the sensitivity of the mCIM for detection of carbapenemase-producing Pseudomonas isolates across 10 sites was found to be 98.0%.12 However in a comparative study by Howard JC et al. the sensitivity of mCIM was found to be only 71.4%.13
Since PCR is the gold standard test for the detection of carbapenemases, we compared the results of mCIM and conventional PCR in a separate group of blaKPC gene-positive and negative isolates of Klebsiella pneumonia. All the 20 blaKPC PCR positive isolates were positive by mCIM, thus giving a sensitivity of 100% for this gene. However, in the case of 15 blaKPC PCR negative isolates, 4(26.6%) were positive which indicates the presence of a carbapenemase other than KPC carbapenemase. Further 11(73.4%) blaKPC PCR negative isolates were negative by this test also because these could have a different resistant mechanism other than carbapenemase production (porin mutations, efflux pumps etc).
CONCLUSION
The mCIM was inexpensive, easy to perform and interpret, supported by the overall excellent reproducibility of the results in Enterobacteriaceae. Comparing with PCR it was found to be an accurate method for the identification of carbapenemase production in Klebsiella Pneumoniae isolates. This test also shows good reproducibility of the results in Pseudomonas isolates. Thus the test aids in the rapid and reliable identification of carbapenemase-producing carbapenem-resistant Enterobacteriaceae in microbiological laboratories and helps in understanding the local epidemiology of resistance to carbapenems, thus serving as one of the important tools to prevent the spread of these drug-resistant pathogens.
Acknowledgement: We thank the staff of the Department of Microbiology, SKIMS Srinagar for their support during the study.
Conflict of interest: The authors declare that the research was conducted in absence of any conflict of interest.
Ethical Clearance: Not Required
Source of funding: No financial support to declare.
Author`s contributions: Author Benazir Shazia designed the study, wrote the protocol and performed the laboratory methods. Author Bhat Asifa wrote the first draft of the manuscript, performed the laboratory methods and the literature searches. Author Fomda BA managed the analysis of the study. All the authors read and approved the final manuscript.
References:
1. Jing X, Zhou H, Min X, Zhang X, Yang Q, Du S et al. The Simplified Carbapenem Inactivation Method (sCIM) for Simple and Accurate Detection of Carbapenemase-Producing Gram-Negative Bacilli. Front. Microbiol. 2018; 9:2391.
2..McMullen AR, Yarbrough ML, Wallace MA, Shupe A, Burnham C-AD. Evaluation of Genotypic and Phenotypic Methods to Detect Carbapenemase Production in Gram-Negative Bacilli. Clin Chem. 2017 Mar;63(3):723–30.
3. Serapsüzükyildiz, Banuka¸ Skatepe, Havvaavciküçük¸ Sükranöztürk. Performance of Carba NP and CIM tests in oxa-48 carbapenemase-producing Enterobacteriaceae. ActaMicrobiologica et Immunologica Hungarica 2017; 64:1,9–16.
4. Pierce VM, Simner PJ, Lonsway DR, Roe-Carpenter DE, Johnson JK, Brasso WB et al. Modified carbapenem inactivation method for phenotypic detection of carbapenemase production among Enterobacteriaceae. J Clin Microbiol. 2017;55:2321–2333.
5. Papagiannitsis CC, Studentova V, Izdebski R, Oikonomou O, Pfeifer Y, Petinaki E, Hrabak J. Matrix-assisted laser desorption ionization-time of flight mass spectrometry meropenem hydrolysis assay with NH4HCO3, a reliable tool for direct detection of carbapenemase activity. J Clin Microbiol 2015;53:1731–1735.
Efficacy of Modified Carbapenem Inactivation Method 10
6. Tamma PD, Opene BNA, Gluck A, Chambers KK, Carroll KC, Simner PJ. A comparison of eleven phenotypic assays for the accurate detection of carbapenemase-producing Enterobacteriaceae. J Clin Microbiol. 2017;55:1046-1055.
7. Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial susceptibility testing.22nd informational supplement (M100-S28). Wayne, PA: CLSI; 2018.
8. Pragasam AK, Veeraraghavan B, Bakthavatchalam YD, Gopi R, Aslam RF. Strengths and limitations of various screening methods for carbapenem-resistant Enterobacteriaceae including new method recommended by clinical and laboratory standards institute: A tertiary care experience. Indian J Med Microbiol 2017;35:116-9.
9. Schechner V, Straus-Robinson K, Schwartz D, Pfeffer I, Tarabeia J, Moskovich R et al. Evaluation of PCR-based testing for surveillance of KPC-producing carbapenem-resistant members of the Enterobacteriaceae family. J Clin Microbiol. 2009 Oct;47(10):3261-5.
10. Tijet N, Patel SN, Melano RG. Detection of carbapenemase activity in Enterobacteriaceae: comparison of the carbapenem inactivation method versus the Carba NP test. J Antimicrob Chemother. 2015;71(1):274–276.
11. Lisboa LF, Turnbull L, Boyd DA, Mulvey MR, Dingle TC. Evaluation of a modified carbapenem inactivation method for detection of carbapenemases in Pseudomonas aeruginosa J Clin Microbiol 2018;56(1):e01234-17.
12. Simner PJ, Johnson JK, Brasso WB, Anderson K, Lonsway DR, Pierce VM et al. Multicenter Evaluation of the Modified Carbapenem Inactivation Method and the Carba NP for Detection of Carbapenemase-Producing Pseudomonas aeruginosa and Acinetobacter baumannii. J Clin Microbiol. 2017 Dec 26;56(1):e01369-17.
13. Howard JC, Creighton J, Ikram R, Werno AM. Comparison of the performance of three variations of the Carbapenem Inactivation Method (CIM, modified CIM [mCIM] and in-house method (iCIM)) for the detection of carbapenemase-producing Enterobacterales and non-fermenters. J. Glob. Antimicrob. Resist. 2020 Jun 1;21:78–82.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License