IJCRR - 13(15), August, 2021
Pages: 154-157
Date of Publication: 10-Aug-2021
Print Article
Download XML Download PDF
An Experimental Study on Classification of Brain Images
Author: Pranati Satapathy, Sarbeswara Hota, Sateesh Kumar Pradhan
Category: Healthcare
Abstract:Introduction: The brain is the major and vital organ of the central nervous system. After the age of 60 or in old age, the human brain may suffer from various disorders. Brain diseases may also occur due to some inevitable causes in the normal human body. As the brain stops functioning, the human body goes into a paralyzed state. To treat various brain diseases, neurologists use different brain imaging techniques. Aims: Magnetic Resonance Imaging (MRI) technique is one of the promising imaging techniques used in recent days for analyzing brain diseases. Manual analysis and classification of brain images into normal or deceased is a tedious task. So different supervised learning techniques are used for this purpose. Methodology: This paper focuses on the experimental study on feature selection using PCA and LDA and classification of two of the brain image datasets i.e. Glioma and Alzheimer. Result: The experimental study suggested that the PCA+MLP classifier obtained accuracy values of 92.68% for the Glioma dataset and 90.023% for the Alzheimer dataset. PLC is used for feature reduction and MLP is used as a classification task. Conclusion: The results suggested that PCA with MLP outperformed the other models.
Keywords: Magnetic Resonance Imaging, Classifier, Perceptron, Decision Tree, Principal Components, Dimension Reduc�tion, Noninvasive
Full Text:
INTRODUCTION
The nervous system is the major constituent of the biological structure of the human body. The vital component of the nervous system is the brain which controls all the operations of the human body. The brain can be affected by several diseases in the life cycle.1 The brain disorders may sometimes lead to death. So it poses a major challenge for neurologists in the treatment of brain diseases. Brain imaging procedures are adopted in this context. Out of the different imaging techniques, the Magnetic Resonance Imaging (MRI) technique is the noninvasive method used in the analysis and identification of brain diseases in recent days. 2, 3
The manual study and interpretation of brain MRI images by medical practitioners is infeasible and may lead to erroneous diagnoses. So the computer-assisted methods are beneficial in this context. Various researchers have used machine learning techniques in classifying the brain images as normal and pathological. 4 The authors applied random forest as the classifier for the brain MRI image classification.5 PCA and neural networks are widely used in the classification of brain images.6,7 The authors adopted Principal Component Analysis (PCA) for feature selection from the extracted brain images.8 They applied Multilayer Perceptron (MLP) for the classification purpose that outperformed k-Nearest Neighbor (k-NN) classifier. Independent Component Analysis (ICA) has also been used as dimension reduction technique. 9, 10 From the literature study, the various steps involved in the brain MRI image classification process can be listed as follows:
-
Brain Image dataset collection
-
Data preprocessing with feature extraction and feature reduction
-
Classification and performance measurement.
The objective of this study is to collect the brain MRI image datasets and then perform data preprocessing. PCA and ICA have been used for dimension reduction purposes. Then three different classifiers i.e. Random forest, k-NN and MLP are applied for the classification purpose. The accuracy values are compared for determining the efficient classifier. This paper is organized as follows. The methodologies are described in the second section. The experimental work is described in the third section. Finally, section 4 discusses the conclusion.
METHODOLOGY
The different feature reduction and classification techniques are discussed in this section.
Independent component analysis (ICA)
ICA is one of the unsupervised dimension reduction strategies used in neuroimaging studies. It groups the original dataset into a set of independent features.11, 12 These independent components are also most relevant to the classification task.
Principal component analysis
PCA is one of the dimension reduction techniques used frequently in the data science and machine learning field. The high-dimensional dataset is reduced to different principal components based on the variance values. 13, 14 No data are lost during the reduction to the low dimensional features. The principal components are the uncorrelated variables and the given initial features are the set of correlated variables.15
Random forest classifier
This is one of the learning approaches used for classification tasks. This is constructed by combining several decision trees. 16 The basic purpose of this ensemble approach is to enhance the training process and classification accuracy. 18 The number of classification trees considered in the random forest approach is chosen randomly.
K-nearest neighbor (k-NN) Classifier
the k-NN classifier is the common classifier used in pattern recognition. The number k is chosen randomly and is very small. The training sample is assigned with the class label that has a minimum distance within the k neighbors. The weights are assigned for different neighbors. It can be stuck at local optima.
Multilayer perceptron (MLP)
MLP is one of the feed-forward neural network structures used for classification purposes. It is completely based on the functionality of the human brain. It belongs to the group of supervised learning. It has one input layer, one or more hidden layers and one output layer. During classification, the number of input neurons will be the number of features. The number of hidden nodes is randomly determined. For the binary classification problems, there is only one output neuron. 16
Proposed Model
The work proposed in this paper is summarized in figure 1. The brain images of the two diseases are collected and preprocessed using 2D-DWT, PCA and ICA. Then the three different classifiers are used for classifying the brain images into normal or diseased. The accuracy values are recorded for the performance measurement ( Figure 1).
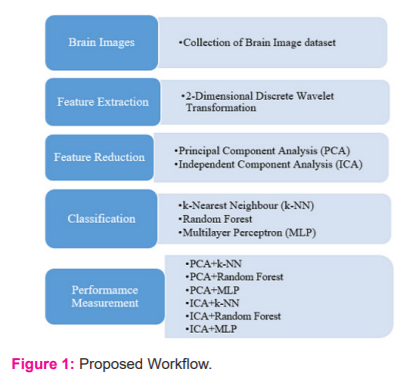
SIMULATION STUDY
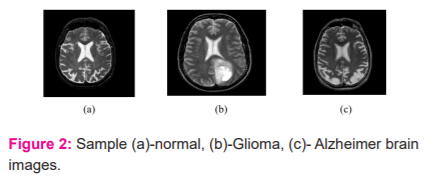
The experimental study is conducted on MRI image datasets of two brain diseases i.e. Glioma and Alzheimer. These two datasets are collected from the Website of Harvard School of medicine. The Glioma dataset contains 122 images and the Alzheimer dataset contains 100 images where each image is of size 288 X 288. The sample MRI images of these two brain diseases are shown in Figure 2-(b) and 2-(c) respectively. Figure 2-(a) shows the normal brain MRI image.
2D DWT is used for feature extraction. Using this procedure, 1296 features are extracted.
FEATURE REDUCTION
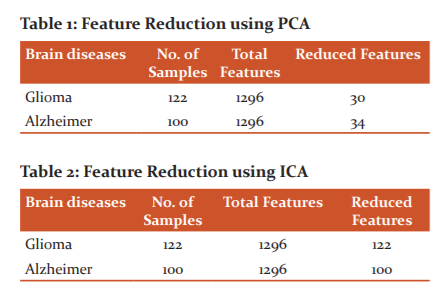
In this work, two feature reduction techniques i.e. PCA and ICA are used. The number of reduced features are shown in Table 1 and Table 2 for PCA and ICA respectively.
CLASSIFICATION
The datasets are divided into train and test datasets for supervised learning. In this work, the Glioma and Alzheimer data sets are divided as 75% for training and 25% for testing purposes. The three types of classifiers i.e. random forest, k-NN and MLP classifiers are considered for the experimental study. 15,16
The loss occurred during training of the datasets using PCA+MLP and ICA+MLP classifiers for the Glioma and Alzheimer datasets are shown in Figure 3 and 4.


The accuracy values found during the classification of the brain diseases are stored in Table 3 and Table 4 for Glioma and Alzheimer image datasets.

From Table 3, it is found that PCA with MLP and PCA with k-NN classifier has same accuracy values i.e. 92.68% for classification of Glioma images into normal and pathological brain.
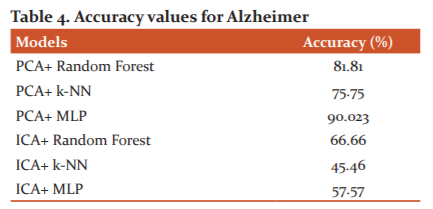

From Table 4, it is found that PCA with MLP classifier has highest accuracy i.e. 90.023% values for classification of Alzheimer images into normal and pathological brains.14,15
CONCLUSION
This work focuses on the experimental study on the performances of different standard classifiers i.e. random forest, k-NN and MLP classifiers for the brain MRI image classification of Glioma and Alzheimer datasets. The 2D DWT is used to extract the features from the brain images. Then two different feature reduction techniques i.e. PCA and ICA are employed in this work to reduce the feature sets. The experimental results suggest that the PCA with MLP produced the highest accuracy values and hence outperformed the other models for the classification of brain images for these two brain diseases.
ACKNOWLEDGEMENT
We acknowledge the immense help received from the scholars whose articles are cited and included in references of this manuscript. We are also grateful to authors / editors / publishers of all those articles, journals and books from where the literature for this article has been reviewed and discussed.
Conflict of Interest
NIL
Source of Funding
NIL
References:
-
Duyn JH. Study of brain anatomy with high-field MRI recent progress,” Magnetic resonance imaging (MRI).2010;28(8):1210–1215.
-
Bauer S, Wiest R, Nolte LP, Reyes M. A survey of MRI-based medical image analysis for brain tumour studies. Phys Med Bio.2013; 58(13):93.
-
Nayak DR, Dash R, Majhi B. Brain MRI image classification using two-dimensional discrete wavelet transform and AdaBoost with random forests. Neurocompt. 2016;177: 188–197.
-
Zhang Y, Dong Z, Wu L, Wang S. A hybrid method for MRI brain image classification. Expt Syst Applic. 2011;38(8):10049–10053.
-
Bartlett M. S., Movellan J. R., Sejnowski T.J., “Face recognition by independent component analysis”, IEEE Transactions on neural networks, 13(6), 1450-1464, 2002.
-
Stone JV. Independent component analysis: an introduction. Tren Cogn Sci. 2002; 6(2): 59-64.
-
Salimi-Khorshidi G. Automatic denoising of functional MRI data: combining independent component analysis and hierarchical fusion of classifiers. Neuroim. 2014; 90: 449-468.
-
Yu SN, Chou KT.Integration of independent component analysis and neural networks for ECG beat classification. Expert Syst Applic.2008; 34(4): 2841-2846.
-
Nandi D., “Principal component analysis in medical image processing: a study”, (IJIM), 1(1), 65-86, 2015.
-
Zhang YD, Wu L. An MR brain images classifier via principal component analysis and kernel support vector machine. Progr Electrom Res. 2012;130:369-388.
-
MingRu K, Zheng Q, Yan SK, Arunkumar N. Medical image classification algorithm based on principal component feature dimensionality reduction”, Future Generation Compt Syst. 2019; 98: 627-634.
-
Khatami A, Khosravi A, Nguyen T, Lim CP, Nahavandi S. Medical image analysis using wavelet transform and deep belief networks. Expt Syst Appl.2017; 86: 190-198.
-
Wei S. Medical image super-resolution by using multi-dictionary and random forest. Sust Citi Soc.2018; 37: 358-370.
-
Ko BC, Kim SH, Nam JY. X-ray image classification using random forests with local wavelet-based CS-local binary patterns. J Digit Imag. 2011;24(6):1141-1151.
-
Aci M, ?nan C, Avci M. A hybrid classification method of k nearest neighbour, Bayesian methods and genetic algorithm. Exp Syst Appl. 2010;37(7):5061-5067.
-
Hassanien AE, Moftah HM, Azar A,Shoman TM. MRI breast cancer diagnosis hybrid approach using adaptive ant-based segmentation and multilayer perceptron neural networks classifier. Appl Soft Comp. 2014;14: 62-71.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License