IJCRR - 13(15), August, 2021
Pages: 106-111
Date of Publication: 10-Aug-2021
Print Article
Download XML Download PDF
Prevalence of HCV Infection and its Genotype among High-Risk Children in Tertiary Care Centre
Author: S. Balasubramanian, D. Selvakumar, D. Nirmala, Sumathi, W. S. Sumathi, T. Pradeebaa
Category: Healthcare
Abstract:Introduction and Objective: Hepatitis C infection (HCV) is one of the important causes of chronic liver diseases (CLD). This causes significant morbidity and mortality in developing countries like India due to its low-level infection control. The destinations of the examination were to investigate parenterally transmitted, HCV in recently analysed haematological disorders, CKD and CLD patients from Madras Medical College and Hospital, Chennai, and further, explore the HCV genotype appropriation in those patients. Materials and Methods: Clinical examination, socio-demographic history, and risk factors exposure and clinical details of each patient were recorded in the proforma. Identification of hostile to HCV antibodies was undertaken using enzyme immunoassay (ELISA KIT). Anti-HCV- seropositive samples were tested for HCV RNA. Hepatitis C virus-ribonucleic acid (RNA) was detected using a Reverse Transcriptase polymerase chain reaction (PCR) assay. HCV genotyping was determined by HCV RT-PCR genotyping test by using the COBAS AMPLICOR LINEAR ARRAY method. Results: Prevention and control of Hepatitis C Virus infection were troublesome as a result of the intricacy present in geological dissemination of HCV and its genotypes, related danger elements and cofactors. HCV type 1 frequently [43/45 = 96% CI (85%- 99%)] more overwhelming in our investigation Conclusion: This study concluded that transfusion of blood and blood products and therapeutic intervention by infected needle are two important preventable modes of spread of hepatitis C virus infection. This made a high risk in children in our country. Improved surveillance and periodic epidemiological studies will have to be undertaken to monitor and prevent this blood-borne virus and its genotype distribution.
Keywords: Chronic hepatitis, Haematological disorders, Genotype, Hepatitis C, Transfusion, Prevalence and chronic kidney disease
Full Text:
INTRODUCTION:
Hepatitis C virus (HCV) is an RNA virus member of the family Flaviviridae.1Approximately 170 million people in the world are infected with HCV. HCV creates an antiviral state, by inhibiting interferon-mediated response. Hence nearly 80% of infections become chronic, among those 20% develop cirrhosis in 5-20 years.
Hepatitis C virus causes continuing inflammation of hepatocytes because of its chronicity leading to CLD, Chronic hepatitis and hepatocellular carcinoma.
Children with chronic disorders like haematological disorders, Chronic Liver disease [CLD], Chronic Kidney disease [CKD] who receives a frequent transfusion of blood, blood products, dialysis and Chemotherapy drugs as a part of their treatment for the above problems are prone to get this blood-borne virus.1
Determining HCV genotype is also important, particularly when therapy is considered because the response to the current therapeutic agents varies greatly. Genotype 1 and genotypes 2, 3 are more reliably responsive to therapy. HCV infection is incessant in patients going through chronic chemo dialysis (HD), with a predominance somewhere in the range of 8 and 10%, and there is a specific concern as a result of the great danger for ongoing liver sickness, inconveniences in renal transplantation, and demise in these patients.2,3
The broad utilization of recombinant erythropoietin to address renal weakness in HD patients brought about a critical decrease in blood bondings. Notwithstanding, past examinations have shown that once more diseases in single HD units may, in any case, happen without other parenteral danger factors.4 It has been proposed that disease could be communicated from one patient to another in the emergency clinic, and there is presently backhanded proof that HCV contamination happens among HD patients during rehashed dialysis methodology, but not through the equipment, probably due to procedural errors. 5Viral titers should be checked yearly to document spontaneous remission. Most patients develop chronic hepatitis.6Progressive liver damage is higher in those with additional comorbid factors such as alcohol consumption, viral genotypic variations, obesity, and underlying genetic predispositions. Referral to a pediatric gastroenterologist is strongly advised to take advantage of up-to-date monitoring regimens and to optimize their enrollment in treatment protocols when available.
With this background, we investigated the proportion of HCV infection in newly diagnosed CLD patients from a Child care institute. In addition, we determined the HCV genotypes and examined their relevance concerning the clinic laboratory presentation. By far most of the patients are tainted with HCV genotypes 1a and 1b. In this examination, we discovered genotype 1a to be the most predominant genotype in the patients contemplated.
Materials and Methods:
This was a hospital-based descriptive study conducted from June 2014 to November 2014 from Madras Medical College and Hospital, Chennai. Ethical approval to offer HCV testing to high-risk children was obtained from the Institutional Ethics Committee, Madras Medical College, Chennai. [Ethical clearance number: 51062014]
Study population:
High-risk children attending the PediatricHematology, Pediatric Nephrology and Pediatric Gastroenterology outpatient department and inpatient wards at the Institute of child health during the study period.
Sample size:
Consecutive samples presenting to the institution during the study period
The sample size was calculated using the formula,

95% confidence interval Zα = 1.96 ≅ 2
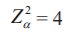
Anticipated error α =5 %
n – Required sample size
– Required sample size
P – Prevalence of Hepatitis c virus infection from previous studies
Q – (1-p)
L – Allowable error
Using the above formula the sample size was calculated as Minimum required sample n= 4× 50×50 /25 =400
High-risk children were examined and assessed for
-
Socio-demographic status including age, sex and residence (urban or rural population),
-
Exposure to risk factors like the previous history of hospitalization, duration of illness, transfusion details, dialysis details, multiple injections (chemotherapy ) and other therapeutic interventions done in the past.
-
Clinical status like Anemia, jaundice, lymphadenopathy, hepatomegaly splenomegaly.
After explaining the procedure, Blood samples of about 3 ml of venous blood were collected under strict aseptic precautions and Serum was separated and tested for the presence of hepatitis B surface antigen (HBsAg). Liver enzymes aminotransferases (SGOT, SGPT) and BILRUBIN were also measured in seropositive cases.
Detection of anti-HCV antibodies was undertaken using third-generation enzyme immunoassay (ELISA KIT) and the tests were performed according to the manufacturer’s instructions provided in the kit.
Anti-HCV- seropositive samples were tested for HCV RNA.
Hepatitis C virus-ribonucleic acid (RNA) was detected using a Reverse Transcriptase polymerase chain reaction (PCR) assay.
HCV genotyping was determined by HCV RT-PCR genotyping test by using the COBAS AMPLICOR LINEAR ARRAY method.
Statistical Analysis
The present study clinical data and laboratory data (ANTI HCV positive - 116, HCV RNA positive -45, GENOTYPE -45) were collected and entered in MS OFFICE EXCEL spreadsheet and the data were analyzed by using the statistical package for social science (SPSS) 17 Version. The results are presented in percentages.
Results:
The present study was undertaken on 435 children during the study period.
Sociodemographic analysis:
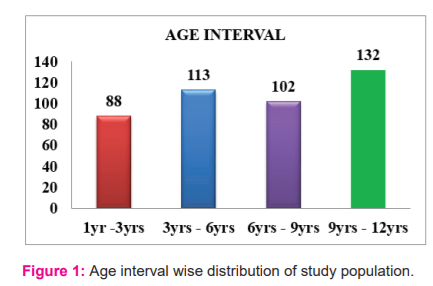
Maximum age 9yrs-12 yrs children constitute 30% of the study population and minimum age 1 yrs -3 yrs children constitute 20% of the study population as shown in figure:1
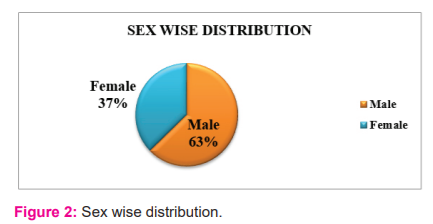
37% of the study population were female sex and 63 % of the study population were male sex as shown in the figure: 2
118 children from an urban area, 317 children from the rural area were included in the study population.
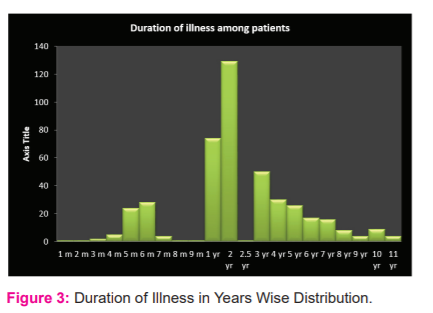
435 children had a history of the previous hospitalization in the study population Figure :3 showing the maximum duration of illness in the population studied was 11 years and the Minimum duration of illness was 1 month
Exposure to risk factors
Table 1 showing 43% of the children had exposure to blood transfusion in the study population and 7.4% of children had been exposed to chemotherapy (Multiple injections)
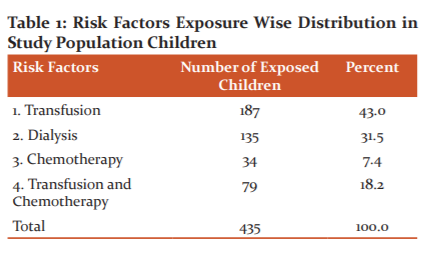
43% of the study population received transfusion, of which maximum number of 30% children received PRBC transfusion and 12% received blood products (FFP, Platelets and Cryoprecipitate)
58% of hematological disorder children, 31% of chronic kidney disease children, 11% of chronic liver disease children were included in this study population as showed in Table 2
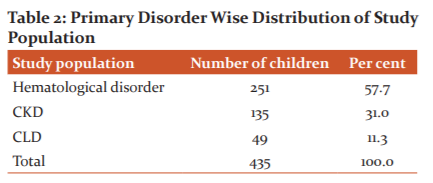

Maximum per cent of the study population were haematological disorder children, of which a maximum of 20% percentage children have acute lymphoid leukaemia and a minimum of 0.2% children have chronic myeloid leukaemia.87 ALL children and 80Thalassemia children and 23 Hodgkin’s Lymphoma children constitute the majority of haematological disorders of the studied population.
Among the total study population, 371 children were anaemic whereas 64 children were normal. Out of 435 children studied 40 children had Lymphadenopathy; Figure: 4 showing that 91% of the study population were normal but 9% had Lymphadenopathy.
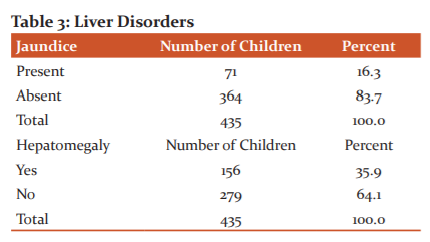
Table 3 shown 71 children were icteric whereas 364 children were anicteric among the total study population. Among the study population, 16% were icteric whereas 84% were anicteric. Out of 435 children, Hepatomegaly was present in 279 children whereas 156 children were normal.
Among the total population studied, splenomegaly was present in 155 children whereas 280 children were normal. Figure: 5 demonstrated that 36% of the total population had splenomegaly; 64% of them were normal.
116 children were seropositive for anti HCV antibodies whereas 319 children were seronegative for Anti HCV antibodies among 435 high-risk children studied as shown in table 4
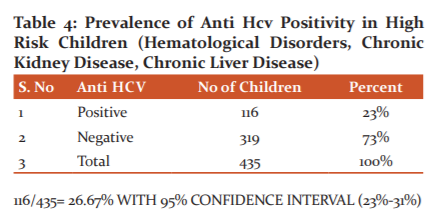

Out of 435 High-risk children attending tertiary care, the hospital studied the seroprevalence of Anti HCV antibodies was 27% whereas 73% of them were seronegative for anti HCV antibodies.
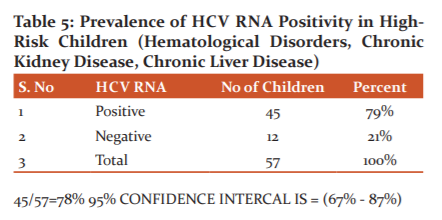
Among 116 seropositive cases, 57 were tested for HCV RNA positivity due to the unavailability of testing kits. The results showed that 45 cases were positive for HCV RNA whereas 12 cases were HCV RNA negative as shown in table 5.
Among 57 children who were positive for HCV RNA 45 children were tested for HCV genotype. Out of 45 children tested a maximum of 43 children had genotype 1, 1 child had genotype 2 and 1 child had genotype 3 as shown in table 6.
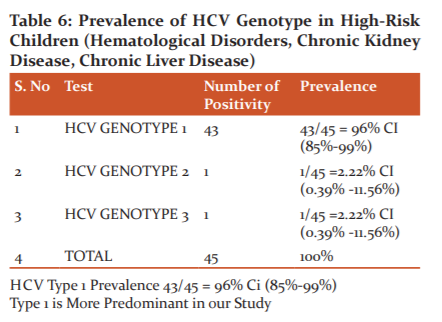
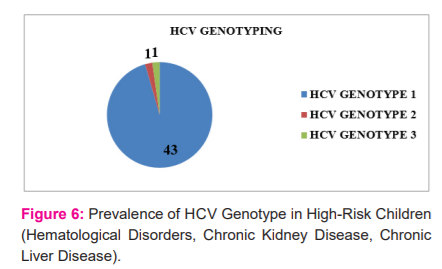
Figure: 6 shows that the prevalence of HCV genotype 1 was 96%, genotype 2 was 1% and genotype 3 was 1% among 45 HCV RNA positive children studied. Hence the most predominant genotype was type 1 in our study involving high-risk children.
Discussion:
In this study prevalence of Hepatitis C virus (HCV) among hospital-based high-risk children is = 26.67%; CI (23%-31%)
The age and sex distribution among the high-risk population are shown in 116 of 435 children who tested positive for antibodies to HCV.
The prevalence among males 25.64% CI (20.82% - 31.13%) and females and 3.3% (95% CI=1.2-5.4%) respectively.
HCV genotype 1 is common in south India which responds well to therapy, whereas genotype 3 predominates are common in the east and west part of India.
There was no statistically significant difference in the exposure rates of males and females (p>0.05). RaghuramanSet al.7conducted a study on the distribution of different genotypes among adult patients attending a tertiary care hospital in south India Christian Medical College and Hospital, Vellore, Tamil Nadu, and they have concluded that HCV genotypes 1 and 3 accounted for 81% of HCV infections in patients from South India region.
J Christmas et al.8 conducted a study of the different genotypes among adults with HCV-related CLD attending a tertiary care hospital in south India during 2002—2012 and concluded that HCV genotype 3(63.85%) and genotype 1 (25.72%), genotypes were detected in their study.
Narahari Set al.9conducted a study on the distribution and prevalence of different genotypes in the Indian patient cohort, in Navi Mumbai, the eastern part of India.
In their study, they have examined and included 2118 patients from different geographic regions of India. Results were analyzed and HCV genotype 3 (3a/3b primarily) in 62% and HCV genotype 1(1a/1b primarily) in 31% of patients were found to be the predominant genotypes in India. They have concluded their study that the predominance of HCV3 was significant in northern (p=0.01) and eastern (p=0.008) regions of India.
Ponamgi SP et al. 10 conducted a study titled Prevalence of hepatitis C virus (HCV) co-infection in HIV infected individuals in South India and characterization of HCV genotypes in a tertiary care hospital in Hyderabad, Andhra Pradesh, India. In their study, they have concluded that the most prevalent genotype in co-infection was genotype 1b.
Anita Chakravarti et al. 11conducted a study on the distribution of different genotypes among adult patients attending various tertiary care hospitals in New Delhi, the northern part of India. HCV Genotype 3 was found to be the most predominant (63%) genotype followed by infection with HCV genotype 1 in 30.98 per cent (22/71) and genotype 2 in 5.63 per cent (4/71) of cases. They have concluded that HCV genotype 3 and 1 accounted for approximately 95% of the HCV infection in Delhi and surrounding areas.
All of the above studies conducted in various parts of India showed HCV genotype 3 followed by genotype 1 were the Predominant Genotypes identified in the adult population whereas Mathavi Chandra et al. study12 showing HCV genotype 1 followed by genotype 3 were the predominant genotype present in south India. Mathavi Chandra et al conducted a study titled Genotyping of Hepatitis C virus (HCV) in infected patients from South India in a tertiary care hospital (Deccan College of Medical Sciences and Allied Hospitals, Kanchanbagh), Hyderabad, and Andhra Pradesh, India. In their study, they have observed that 60% of the patients (30 patients with chronic hepatitis) were infected with HCV genotype 1 and 40% of the patients (4 chronic hepatitis patients, 12 patients with chronic renal failure and 4 cirrhosis) were infected with genotype 3 of HCV.
They concluded that predominant genotypes of HCV in South India include types 1 and 3. In our study, the most predominant genotype present is genotype 1 followed by genotypes 2 & 3. Our study is conducted in high-risk children with haematological disorders, chronic kidney disease and chronic liver disease.
Prevalence of HCV genotype in high-risk children (haematological disorders, chronic kidney disease, chronic liver disease) HCV type 1 prevalence 43/45 = 96% ci (85%-99%). Type 1 is more predominant in our study
Our study finding shows the most predominant genotype is genotype 1 which is similar to Mathavi Chandra et al. 12 studies which are Conducted on adult population in south India, Andhra Pradesh
Conclusion:
This study concluded that unsafe transfusion of blood and blood products and unsafe therapeutic intervention by infected needle are two important preventable modes of spread of hepatitis c virus infection in at-risk children in our country.
Prevention and control of Hepatitis C Virus infection are difficult because of the complexity present in the geographical distribution of HCV and its genotypes, associated risk factors and cofactors.
Improved surveillance and periodic epidemiological studies will have to be undertaken to monitor and prevent this blood-borne virus and its genotype distribution.
ACKNOWLEDGEMENT:
This research was supported/partially supported by Madras medical college and hospital. We are thankful to our colleagues who provided expertise that greatly assisted the research.
AUTHORS CONTRIBUTION:
S.Balasubramaniamand D.Selvakumarmade substantial contributions to conception, acquisition of data, took part in drafting the article or revising it critically for important intellectual content, D Nirmala, Dr. Sumathi, WS Sumathi and T Pradeepamade Statistical analysis and final approval of the version to be published, and agreed to be accountable for all aspects of the work
SOURCE OF FUNDING: No funding sources
CONFLICT OF INTEREST: None declared
ETHICAL APPROVAL: The study was approved by the Institutional Ethics Committee and the IEC letter number is MMC/IEC- 321/2020
References:
-
Lavanchy D. The global burden of hepatitis C. Liver Int. 2009; 29(1):74-81.
-
Ghany MG. Diagnosis, management, and treatment of hepatitis C: An update. Hepatology. 2009; 49(4): 1335-74.
-
Choo QL. Isolation of a cDNA clone derived from a blood-borne non-A, non-B hepatitis genome. Sci. 1989; 244(4902): 359-62.
-
Tokita H et al. Hepatitis C virus variants from Thailand are classifiable into five novel genotypes in the sixth (6b), seventh (7c, 7d) and ninth (9b, 9c) major genetic groups. J. Gen. Virol 1995; 76(Pt 9): 2329-35.
-
Sievert W et al. A systematic review of hepatitis C virus epidemiology in Asia, Australia and Egypt. Liver Int. 2011; 31(2):61–80.
-
Zein NN. Clinical significance of hepatitis C virus genotypes. Clin Microbiol Rev. 2000; 13(2): 223-35.
-
Raghuraman S. Distribution of the different genotypes of HCV among patients attending a tertiary care hospital in south India. J Clin Virol. 2003; 26: 61-9.
-
Christdas J, Sivakumar J, David J, et al. Genotypes of hepatitis C virus in the Indian sub-continent: a decadelong experience from a tertiary care hospital in South India. Indian J Med Microbiol. 2013;31(4):349-53.
-
Narahari S, Juwle A, Basak S and Saranath D. Prevalence and geographic distribution of Hepatitis C Virus genotypes in Indian patient cohort. Infect, Gen J. 2009; 9(4):643-645.
-
Ponamgi SP. Prevalence of hepatitis C virus (HCV) co-infection in HIV infected individuals in South India and characterization of HCV genotypes. Indian J Med Microbiol. 2009; 27(1):12-16.
-
Anita Chakravarti. Distribution pattern of HCV genotypes & its association with viral load. Indian J Med Res. March 2011; 2: 326-331.
-
Madhavi Chandra. Prevalence of hepatitis C virus (HCV) genotypes among positive south indian patients. Molec Biol Rep. 2011; 38(4):2719–22.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License