IJCRR - 13(15), August, 2021
Pages: 51-57
Date of Publication: 10-Aug-2021
Print Article
Download XML Download PDF
A Review on Esophageal Cancer Detection and Classification Using Deep Learning Techniques
Author: Kumar C A, Mubarak M N D
Category: Healthcare
Abstract:Introduction: Esophageal cancer (EC)is the sixth most common cancer with a high fatality rate. Early prognosis can improve the survival rate of the patients. The sequence of the progress of the EC is from Esophagitis to Non-Dysplasia Barrett's Esophagus to Dysplasia Barrett's Esophagus to Esophageal Adenocarcinoma (EAC). Computer-Aided Diagnosis (CAD) has become a primary tool of the decade to diagnose various diseases. Objective: The recent advances in Artificial Intelligence (AI), Machine Learning (ML), and Deep Learning (DL) have enriched the potential of detection, localization, and classification of the medical image pattern. Here, we have compiled multiple research works based on supervised DL architectures. Methods: This review focuses on the application of DL techniques for the detection, segmentation, and classification of the various stages leading to the EAC. The surveyor concentrates on the pre-trained classification detection and segmentation models. Results: The advancements in AI have enhanced the contributions in the medical field applications. The technological progress in AI and DL led to a large number of researches in the medical field. The new algorithms and DL models resulted in many automated systems for the detection segmentation and classification of oesophagal cancer. Conclusion: This review discusses the various challenges, limitations, and future aspects of analysing endoscopic images based on DL methods. Further investigations are to be carried out to improve the performance of CAD systems for successful real-time detection of oesophagal and associated stages. It is essential to formulate more collaborated studies with experts in the field.
Keywords: Barrett’s Esophagus, Computer-Aided Diagnosis, Convolution Neural Networks, Deep Learning, Esophageal Adenocarcinoma, Machine Learning
Full Text:
INTRODUCTION
Acid reflux disease (GERD) is a condition that causes acidity and indigestion.1 The stomach acid's backwash will gradually replace the healthy oesophagal tissue with the tissue that resembles the intestinal/gastric tissues. Figure:1 shows the EC progression as a continuum variation from Esophagitis to Non-Dysplasia Barrett's Esophagus to Dysplasia Barrett's Esophagus to EAC. The change in the texture of the oesophagal tissue lining in Barrett’s Esophagus (BE)may lead to dysplasia. Based on the variation in the texture pattern of the oesophagus lining, the abnormal tissue cells can be categorized as non-dysplastic (noncancerous), High-grade dysplasia (HGD), and Low-grade dysplasia (LGD).2 The condition of BE with High-grade dysplasia is an advanced pre-cancer stage leading to EAC. and EAC's occurrence has drastically increased in Western countries, especially in industrial countries. One of the main reasons for the drastic increase in the rate is due to society's unhealthy lifestyles. The sixth most common cancer with a high fatality rate is EC3. An early prognosis can improve the survival rate of the patient ( Fig 1).
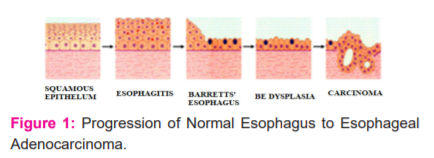
CAD is one of the most prominent tools for providing an improved prognosis for EC. In recent years AI has made remarkable progress in the medical domain. The deep neural network's ability to automatically learn significant low-level features and combine them with high-level features enhances DL performance in medical applications.4 DL with CNN is one of the most efficient and standard tools to perform multiple tasks like object recognition, classification, and segmentation.3-6 We aim to review and study the feasibility of DL for a CAD system using CNN for the prognosis of premalignant and malignant stages of BE (Fig 2).
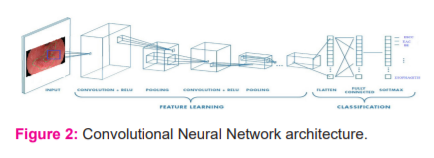
DEEP LEARNING MODELS
AI is an area of rapid technological advancement, especially in medical applications. AI equips the computer to emulate human intelligence5. Both Machine Learning (ML) and DL are a subset of AI. The DL technique addressed all the major concepts involved in image processing.6 The DL structure has deep CNN networks with a significant number of hidden layers as shown in Figure:2. In health care applications, the DL is considered state-of-the-art technology by its performance.7 Figure:3 depicts the supervised DL models for medical image processing.
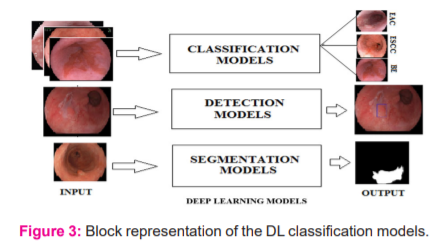
Classification Models
The input image traverse through a set of convolutional kernels with down-sampling. A softmax layer towards the end of the model will generate the class probability for classification.8 The model performs the classification process based on the category, subcategory, lesion-based analysis, and lesion morphology like invasion depth. The various classification models are AlexNet,9 VGG,10 GoogLeNet,11 Inception V4,12 Residual Network (ResNet)13and DenseNet.14
Segmentation Models
The segmentation process is the delineation of the lesions' borders under consideration15. Generally, all the segmentation models have two modules, an encoder and a decoder8. The encoder performs the convolution of the image with kernels and downsamples the feature maps. The decoder module performs the deconvolution and the up-sampling process. The model performs both the Semantic and Instance segmentation. Some of the popular segmentation models are Fully Convolution Network (FCN),16 SegNet,17 U-Net,18 Deep Lab,19 DeepLabV3.20
Detection Models
The complete knowledge of an image is conceived when we accurately understand the object's nature and location in the picture.21 The object detection model needs to establish the object location (localization) and the category of the object(classification). A deeper analysis needs to be done by the model to predict the subcategory of the objects. The detection will be based on the object scores obtained from the feature extracted by the fully connected layers in some models. Multiple object detection can also be performed based on the class and object score derived from the feature extracted. The R-CNN,22 Fast R-CNN,23 Faster R-CNN,24 Mask R-CNN,25 Single Shot Multi-BoxDetector (SSD)26 and YOLO27are some of the prevalent detection models used.
SURVEYED WORKS
Takiyama et al.28 propose a GoogLeNet model trained by a back-propagation algorithm for the organ's anatomical classification, larynx, oesophagus, stomach, and duodenum. The network is fine-tuned with an ADAM optimizer with a learning rate of 0.0002.
Kumagai et al.29proposes a DL model for the classification of ESCC. Endocytoscopy(ECS) is a magnified endoscopic method that enables the observation of surface epithelial cells in real-time. The ECS enables a real-time optical biopsy. The ECS performs tissue analysis and perceives the histological features in real-time. The diagnosis can be made without performing biopsy reference of the histology using GoogLeNet trained by a back-propagation algorithm. Ada Delta fine-tunes the training of the model with a learning rate of 0.004.
In Tokai et al.30 the ESCC detection is performed with SSD at a rate of 95.7% in 10sec. The GoogLeNet evaluated the measure of invasion depth of ESCC and the sub-classification for WLI and NBI. The network was trained with all the CNN layers fine-tuned by stochastic gradient descent (SGDM) with a learning rate of 0.0001.
Van der Putten et al.31 proposes a combination of multiple modalities to analyze early neoplasia in BE for better localization accuracy. The sweet spot and soft spot is predefined, and the image registration technique is applied for aligning BLI to WLI. The Canny edge detection is applied for enhancing image pairs.Resnet18 is used for the classification of the patches, which is fine-tuned with an ADAM optimizer.
Tomita et al.32proposes an attention-based model with Resnet18 for the tissue level classification of the histological image into normal, BE with no dysplasia, BE with dysplasia, and EAC. Grid-based feature extraction performed with Resnet18, followed by a 3D CNN kernel to build an attention map. The attention map with the attention feature weights is combined to obtain the feature vector for the tissue classification. The model is trained with high-resolution images and fine-tuned with an ADAM optimizer.
Liu et al. 33proposes an Inception- ResNet model to classify premalignant cancer and EC lesions. The original and preprocessed image is applied as input to the Inception -ResNet model separately for feature extraction. The features obtained are concatenated through a concatenation fusion function. The model achieved better performance when trained and fine-tuned with SGDM optimization with a learning rate of 0.001.
Ebigbo et al. 34 proposed a ResNet network for BE analysis. The pathologically validated images served as a reference for classification. The images labelled by experts serve as the standard of reference for the segmentation process. All validation was carried out using Leave-One-Patient-Out Cross-validation (LOpOCV). The small patches are extracted from the endoscopic colour images, and augmentation is applied to obtain diversified images of a similar class. The classification of the full image was attained by culminating the probability of each patch class.
Hong et al.35proposes a simple CNN architecture to differentiate between sub-classes IM, GM, and NPL. The proposed CNN yielded 80.77 per cent classification accuracy among sub-classes of IM, GM, and NPL.
Van Riel et al.36proposes a new technique for achieving real-time performance in early EC diagnosis using CNN and transfer learning. Performance analysis of some of the popular pre-trained networks such as AlexNet, VGG'16, and GoogLeNet was evaluated using the knowledge transferred from the ImageNet dataset with classifiers such as Support Vector Machine and Random Forest individually. The window-based approach out-performed the existing methods and achieved an AUC of 0.92 (area under the curve).
Mendel et al.37proposes an automated CNN model for the early diagnosis of EAC from High-definition endoscopic images using transfer learning. The ResNet architecture was used to train the models and leave one patient out cross-validation (LOPO-CV) method for evaluation.
Rezvy et al.38focus on a modified Mask-RCNN to detect and segment the Precancerous, BE, High-GradeDysplasia (HGD), EAC and Polyps trained with feature representation in transfer learning mode. The network head of ResNet101 trained on the COCO dataset is used to replace the network head of the AI model. The new model is acquainted with the augmented dataset and fine-tuned with ADAM optimization with a learning rate of 0.0001 for the detection.
Gao et al.39 explore the viability of attaining synchronized processing of endoscopic video for determining the pre-cancer status of squamous cell cancer. The detection architectures, mask-RCNN, and YOLOv3 are used to detect and segment endoscopic images. The images are classified as SCC's 'cancer,' 'high risk,' and 'suspicious.' The Mask RCNN outperformed YOLOv3 both in generating the masks and classification accuracy. YOLOV3 processed the video frames at a rate of 0.1spf, which is ten times faster than Mask RCNN.
Hori et al.40 proposed an AI-based diagnostic system with SSD for diagnosing EAC and Squamous Cell Carcinoma (SCC). CNN precisely diagnosed all the cases of ECs from the combined analysis of WLI and NBI images. The diagnostic system was able to recognize even the smallest lesions (<10mm). CNN diagnostic model achieved a more enhanced performance with the comprehensive diagnosis of NBI and WLI images.
In Ghatwary et al.41, the study focuses on the evaluation of EAC regions from the high-definition white-light endoscopy (HD-WLE) images by using different DLobject detection methods. The SSD, Region-based Convolution Neural Network (R-CNN), Faster R-CNN, Fast R-CNN models explored in this study. From the experimental results, we can infer that the SSD outclass all other approaches. The pre-trained model VGG16 was used for better classification of the EAC.
Zhao et al.42proposes a double-labelled FCN model with multi-task learning for the detection and semantic segmentation of ESCC. The FCN, when the trained end to end within the self-transfer learning framework, optimizes the Region of Interest and Semantic Labels simultaneously. The double labelling FCN model can provide extra attributes for better pixel classification. The type A classification is made for those with inflammations. Type B1 and B2 show the invasion depth of ESSC, which can be treated by local resection, and type B3, the invasion depth is >200µm, which requires surgical treatment. The double-labelled FCN model has a better performance with improved diagnostic accuracy.
Liu et al.43proposes a DeepLabV3+ model for early EC detection and segmentation. DeepLabV3+ has an Xception architecture with Atrous Convolution (AC) and Atrous Spatial Pyramidal Pooling(ASPP). The encoder stage extracts in-depth features by applying AC, and then the feature map is applied to ASPP for generating multiscale features. The decoder stage refines the segmentation process. The feature from the AC in the encoder is simplified via a 1x1 CNN and concatenated with multiscale features. Again, these features are applied to a 3x3 CNN and up-sampled to generate a binary semantic segmentation. The binary segmented image is further processed using the morphological and hole filling process to obtain the annotated image.
Guo et al.44 explore a CAD SegNet model for real-time identification and segmentation of pre-cancerous lesions and early ESCC. SegNet is a deep encoder-decoder module for multiclass segmentation. The encoders extract the low-resolution features with the boundary information stored in the max-pooling indices. The decoders consist of a pixel-wise classifier. It up-samples the attribute maps using the max-pooling indices of the analogous encoder for generating sparse attribute maps. The up-sampling feature map obtained is then convolved to achieve the dense feature map. The dense feature map will reduce the number of parameters, so SegNet needs less memory space and requires less computational time.
Van der Putten. et al.45AI proposes a network Gastro Net model with multi-task learning is proposed to obtain better localization, classification, and semantic segmentation of the BE. A ResNet replaces the CNN network in the encoder and decoder path of the U-Net. The fully connected feature layer and classification layer is added to the bridge network to multi-task learning. Both the classification and segmentation processes perform simultaneously in a single training. Pseudo Labeling, a semi-supervised learning algorithm with Bootstrapping and Ensemble learning, provides more suitable descriptive attributes for enabling multi-stage transfer learning. The model is trained and fine-tuned with ADAM and AMS grad with a weight decay of 10-5 with a cosine cyclic learning rate schedule.
Omura et al.46 focus on a DL model for early EAC detection using a four-layer neural network with the feature extraction performed by Dyadic Wavelet Transform (DYDWT). The input RGB image is converted into HSV and CIELa*b*. A fusion image is constructed by normalizing the S, a*b* components. This fusion image undergoes a contrast enhancement to obtain Sa*bhist*. The new image undergoes a 3-level decomposition using DYDWT. The Inverse DYDWT is applied for image reconstruction. The reconstructed image is the input to the neural network for classification. The DYDWT reduces the input features resulting in faster learning and high computational speed.
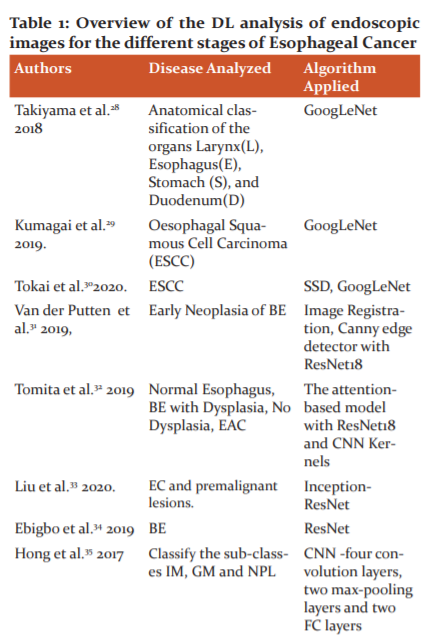
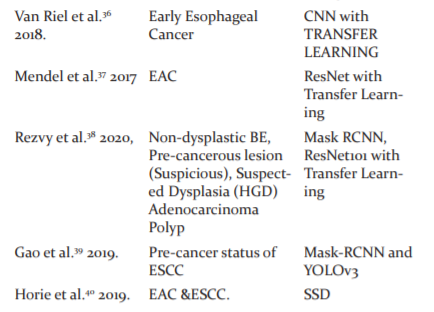
Ghatwary et al.47proposed a DL method for the automatic detection and classification of oesophagal abnormalities. The local features extracted through the Gabor filters and the CNN(DenseNet) are concatenated for enhancing the detection of abnormal regions using Faster RCNN. The overview of the DL analysis of endoscopic images for the different stages of Esophageal Cancer is shown in Table :1.


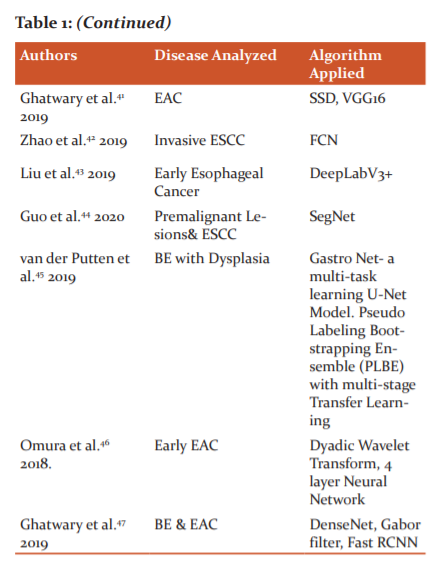
DISCUSSIONS
The technological progress in AI and DL led to a large number of research in the medical field. The new algorithms and DL models resulted in many automated systems for the detection, segmentation, and classification of EC. CNN is currently the backbone of all the DLarchitectures.
Many new network models are based on the multi-dimensional arrangements of CNN layers. Most of the currently used DL models are supervised learning type models. The researchers need to work on the semi-supervised and unsupervised models for the diversified analysis of EC. Studies need to be done on the combination of tasks and networks for better feature extraction. One of the significant drawbacks of medical image analysis is the lack of availability of medical data. The labelling or annotation of the available dataset using experts is a more challenging task. The data augmentation and annotation methods need to be experimented with to combat these challenges. The researchers must have a substantial public dataset of medical images only. A pre-trained network trained in the medical dataset provides more relevant attributes and better diagnostic accuracy through transfer learning.
CONCLUSION
The advancements in artificial intelligence and DL have contributed a lot to develop many CAD techniques for detecting EC in the early stage itself. This review concentrates on some of the recent studies on the diagnosis of different EAC stages with DLtechniques using CNN. More research works are to be carried out with the semi-supervised and unsupervised learning methods for the widespread analysis of the EC. With the invention of the deep generative models and application of hybrid networks, more evaluation and estimation of early oesophagal cancer features can be performed. One of the significant limitations is the lack of availability of data set in the medical fields. The DL approach using CNN and the transfer learning techniques can resolve this problem. Data augmentation seems to be a suitable solution to this unbalanced data availability. The development of many unsupervised network architectures is to be initiated in the future to overcome the dataset availability.
ACKNOWLEDGMENT
The authors acknowledge the immense help received from the scholars whose articles are cited and included in references to this manuscript. The authors are also grateful to authors/ editors/publishers of all those articles, journals, and books from which the literature for this article has been reviewed and discussed.
Conflict of Interest
The authors have no competing conflict of interest to declare.
Source of Funding
There is No funding obtained from any organization for carrying out this research work.
Author Contribution
CHEMPAK KUMAR. A - Conceptualization, Formal analysis, Visualization, Writing -original draft, Writing
-review & editing.
D.MUHAMMAD NOORUL MUBARAK- Writing -review & editing.
References:
-
https://wwwmayoclinic.org[Online]. [14August2019] Available from: https:// www. mayoclinic.org/diseases-conditions/ esophagitis / diagnosis-treatment/drc-20361264
-
www. cancer. org [Online] [4 August 2020] Available from: https: // www.cancer.org/cancer/esophagus-cancer/causes-risks-prevention/risk-factors.html
-
Pakzad R, Mohammadian-Hafshejani A, Khosravi B, Soltani S, Pakzad I, Mohammadian M, Salehiniya H, Momenimovahed Z. The incidence and mortality of oesophagal cancer and their relationship to development in Asia. Annals of translational medicine. IEEE Access. 2017: 9375-9389.
-
Fourcade A, Khonsari RH. Deep learning in medical image analysis: A third eye for doctors. Journal of stomatology, oral and maxillofacial surgery. 2019 Sep 1;120(4):279-88.
-
Hatt M, Parmar C, Qi J, El Naqa I. Machine (deep) learning methods for image processing and radiomics. IEEE Transactions on Radiation and Plasma Medical Sciences. 2019 Mar 1;3(2):104-8.
-
Du W, Rao N, Liu D, Jiang H, Luo C, Li Z, Gan T, Zeng B. Review on the applications of deep learning in the analysis of gastrointestinal endoscopy images. Ieee Access. 2019; 30(7):142053-69.
-
Alom MZ, Taha TM, Yakopcic C, Westberg S, Sidike P, Nasrin MS, Hasan M, Van Essen BC, Awwal AA, Asari VK. A state-of-the-art survey on deep learning theory and architectures. Electronics. 2019 Mar;8(3):292.Krizhevsky, Alex, Ilya Sutskever, and Geoffrey E. Hinton. "Imagenet classification with deep convolutional neural networks." In Advances in neural information processing systems.2012; 1097-1105.
-
Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. 2019; 9(4): 231.
-
Szegedy C, Liu W, Jia Y, Sermanet P, Reed S, Anguelov D, Erhan D, Vanhoucke V, Rabinovich A. Going deeper with convolutions. InProceedings of the IEEE conference on computer vision and pattern recognition 2015; 1-9.
-
Szegedy C, Ioffe S, Vanhoucke V, Alemi A. Inception-v4, inception-resnet and the impact of residual connections on learning. InProceedings of the AAAI Conference on Artificial Intelligence. 2017; 02(12):321.
-
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. InProceedings of the IEEE conference on computer vision and pattern recognition. 2016;770-778.
-
Huang G, Liu Z, Van Der Maaten L, Weinberger KQ. Densely connected convolutional networks. InProceedings of the IEEE conference on computer vision and pattern recognition. 2017;4700-4708.
-
Ebigbo A, Palm C, Probst A, Mendel R, Manzeneder J, Prinz F, de Souza LA, Papa JP, Siersema P, Messmann H. A technical review of artificial intelligence as applied to gastrointestinal endoscopy: clarifying the terminology. Endoscopy international open. 2019 Dec;7(12):E1616.
-
Long J, Shelhamer E, Darrell T. Fully convolutional networks for semantic segmentation. InProceedings of the IEEE conference on computer vision and pattern recognition 2015;3431-3440.
-
Badrinarayanan V, Kendall A, Cipolla R. Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE transactions on pattern analysis and machine intelligence. 2017 Jan 2;39(12):2481-95.
-
Ronneberger O, Fischer P, Brox T. U-net: Convolutional networks for biomedical image segmentation. InInternational Conference on Medical image computing and computer-assisted intervention. 2015; 5 :234-241. Springer, Cham.
-
Chen LC, Papandreou G, Kokkinos I, Murphy K, Yuille AL. Deeplab: Semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected crfs. IEEE transactions on pattern analysis and machine intelligence. 2017 Apr 27;40(4):834-48.
-
Chen LC, Zhu Y, Papandreou G, Schroff F, Adam H. Encoder-decoder with atrous separable convolution for semantic image segmentation. InProceedings of the European conference on computer vision (ECCV). 2018; 801-818.
-
Zhao ZQ, Zheng P, Xu ST, Wu X. Object detection with deep learning: A review. IEEE transactions on neural networks and learning systems. 2019 Jan 28;30(11):3212-32.
-
Girshick R, Donahue J, Darrell T, Malik J. Rich feature hierarchies for accurate object detection and semantic segmentation. InProceedings of the IEEE conference on computer vision and pattern recognition. 2014;580-587.
-
Girshick R, Fast RC. IEEE Int. Conf. Comput. Vis. Santiago, Chile, December. 2015:7-13.
-
Ren S, He K, Girshick R, Sun J. Faster R-CNN: towards real-time object detection with region proposal networks. IEEE transactions on pattern analysis and machine intelligence. 2016 Jun 6;39(6):1137-49.
-
He K, Gkioxari G, Dollár P, Girshick R. Mask r-cnn. InProceedings of the IEEE international conference on computer vision 2017; 2961-2969.
-
Liu W, Anguelov D, Erhan D, Szegedy C, Reed S, Fu CY, Berg AC. SSD: Single shot multibox detector. In European conference on computer vision 2016;10(8):21-37.
-
Redmon J, Divvala S, Girshick R, Farhadi A. You only look once: Unified, real-time object detection. InProceedings of the IEEE conference on computer vision and pattern recognition 2016;779-788.
-
Takiyama H, Ozawa T, Ishihara S, Fujishiro M, Shichijo S, Nomura S, Miura M, Tada T. Automatic anatomical classification of esophagogastroduodenoscopy images using deep convolutional neural networks. Scientific reports. 2018 May 14;8(1):1-8.
-
Kumagai Y, Takubo K, Kawada K, Aoyama K, Endo Y, Ozawa T, Hirasawa T, Yoshio T, Ishihara S, Fujishiro M, Tamaru JI. Diagnosis using deep-learning artificial intelligence based on the end cystoscopic observation of the oesophagus. Oesophagus. 2019 Apr;16(2):180-7.
-
Hirasawa T, Aoyama K, Tanimoto T, Ishihara S, Shichijo S, Ozawa T, Ohnishi T, Fujishiro M, Matsuo K, Fujisaki J, Tada T. Application of artificial intelligence using a convolutional neural network for detecting gastric cancer in endoscopic images. Gastr Canc. 2018 Jul;21(4):653-60.
-
van der Putten J, Wildeboer R, de Groof J, van Sloun R, Struyvenberg M, van der Sommen F, Zinger S, Curvers W, Schoon E, Bergman J, de With PH. Deep learning biopsy marking of early neoplasia in Barrett’s oesophagus by combining WLE and BLI modalities. In2019 IEEE 16th International Symposium on Biomedical Imaging. 2019; 4(8):1127-1131). IEEE.
-
Tomita N, Abdollahi B, Wei J, Ren B, Suriawinata A, Hassanpour S. Attention-based deep neural networks for detection of cancerous and precancerous oesophagus tissue on histopathological slides. J Ame Mat Netw Open. 2019 Nov 1;2(11):e1914645-.
-
Liu G, Hua J, Wu Z, Meng T, Sun M, Huang P, He X, Sun W, Li X, Chen Y. Automatic classification of oesophagal lesions in endoscopic images using a convolutional neural network. Annal Transl Med. 2020 Apr;8(7).
-
Ebigbo A, Mendel R, Probst A, Manzeneder J, de Souza Jr LA, Papa JP, Palm C, Messmann H. Computer-aided diagnosis using deep learning in the evaluation of early oesophageal adenocarcinoma. Gut. 2019 Jul 1;68(7):1143-5.
-
Hong J, Park BY, Park H. Convolutional neural network classifier for distinguishing Barrett's oesophagus and neoplasia endomicroscopy images. In2017 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) 2017;7( 11): 2892-2895.
-
Van Riel S, Van Der Sommen F, Zinger S, Schoon EJ, de With PH. Automatic detection of early oesophagal cancer with CNNS using transfer learning. In 2018 25th IEEE International Conference on Image Processing (ICIP) 2018; 7: 1383-1387. IEEE.
-
Mendel R, Ebigbo A, Probst A, Messmann H, Palm C. Barrett’s esophagus analysis using convolutional neural networks. InBildverarbeitung für die Medizin. 2017 ;80-85. Springer Vieweg, Berlin, Heidelberg.
-
Rezvy S, Zebin T, Braden B, Pang W, Taylor S, Gao X. Transfer learning for Endoscopy disease detection and segmentation with mask-RCNN benchmark architecture. In2020 IEEE 17th International Symposium on Biomedical Imaging 2020;17.
-
Gao X, Braden B, Taylor S, Pang W. Towards real-time detection of squamous pre-cancers from oesophageal endoscopic videos. In2019 18th IEEE International Conference On Machine Learning And Applications (ICMLA) 2019;12(16):1606-1612.
-
Horie Y, Yoshio T, Aoyama K, Yoshimizu S, Horiuchi Y, Ishiyama A, Hirasawa T, Tsuchida T, Ozawa T, Ishihara S, Kumagai Y. Diagnostic outcomes of oesophagal cancer by artificial intelligence using convolutional neural networks. Gastroint Endsc. 2019 Jan 1;89(1):25-32.
-
Ghatwary N, Zolgharni M, Ye X. Early oesophagal adenocarcinoma detection using deep learning methods. Int J Comp Assi Rad Surg. 2019 Apr;14(4):611-21.
-
Zhao YY, Xue DX, Wang YL, Zhang R, Sun B, Cai YP, Feng H, Cai Y, Xu JM. Computer-assisted diagnosis of early esophageal squamous cell carcinoma using narrow-band imaging magnifying endoscopy. Endoscopy. 2019 Apr 1;51(04):333-41.
-
Liu DY, Jiang HX, Rao NN, Luo CS, Du WJ, Li ZW, Gan T. Computer-aided annotation of early esophageal cancer in gastroscopic images based on deeplabv3+ network. InProceedings of the 2019 4th International Conference on Biomedical Signal and Image Processing (ICBIP 2019) 2019; 8 (13):56-61.
-
Guo L, Xiao X, Wu C, Zeng X, Zhang Y, Du J, Bai S, Xie J, Zhang Z, Li Y, Wang X. Real-time automated diagnosis of precancerous lesions and early oesophagal squamous cell carcinoma using a deep learning model (with videos). Gastroint Endosc. 2020 Jan 1;91(1):41-51.
-
van der Putten J, de Groot J, van der Sommen F, Struyvenberg M, Zinger S, Curvers W, Schoon E, Bergman J. Pseudo-labeled bootstrapping and multi-stage transfer learning for the classification and localization of dysplasia in Barrett’s oesophagus. International Workshop on Machine Learning in Medical Imaging 2019;10(13): 169-177. Springer, Cham.
-
Omura H, Minamoto T. Detection Method of Early Esophageal Cancer from Endoscopic Image Using Dyadic Wavelet Transform and Four-Layer Neural Network. information Technology-New Generations 2018: 595-601. Springer, Cham.
-
Ghatwary N, Ye X, Zolgharni M. Esophageal abnormality detection using descent-based faster r-CNN with Gabor features. IEEE Access. 2019 Jun 27;7:84374-85.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License