IJCRR - 13(12), June, 2021
Pages: 210-214
Date of Publication: 22-Jun-2021
Print Article
Download XML Download PDF
3D Densealexnet Model for Brain Tumour Segmentation
Author: M. Sumithra, S. Malathi
Category: Healthcare
Abstract:Background: The collection of anomalous cells within or around the brain is stated as a brain tumour. Automatic brain tumour segmentation is considered a challenging task due to complexity and gradient diffusion. To improve the segmentation of 3D brain MRI images Deep Neural Network (DNN) is evolved. However, it is subjected to the drawback of training computational power and complexity. Objective: In this paper, proposed a 3D Dense AlexNet model with backpropagation for segmentation of tumour in brain MRI images. The developed architecture consists of Neural Network for processing input 3D images. This paper focused on improving the overall segmentation process with the Alexnet model for 3D brain images for performance improvement. Method: Based on the training and validation test self-constrained 3D Dense ALexNet model is developed. Within the 3D Dense AlexNet backpropagation is adopted for removing complexity in the testing process and accuracy improvement. Based on the training and testing process 3D MRI image sequences are trained and processed for segmentation on the tumour. Result: The analysis of results expressed that the proposed 3D Dense AlexNet exhibits improved segmentation performance. Based on the proposed 3D AlexNet architecture MRI images are segmented with minimal time. The performance of the proposed 3D Dense AlexNet model exhibited the improved accuracy of tumour detection with reduced computational complexity
Keywords: 3D Brain MRI, Dense AlexNet, Back Propagation, Segmentation, Deep Neural Network (DNN), Neural Network
Full Text:
A tumour is a collection or mass of abnormal cells that occur in various parts of the body. A tumour can result in cancer, which is the main reason for death and accounts for around 13% of every death worldwide. The cancer occurrence rate is rising at an alarming rate in the world. Therefore, tumour detection is significant in previous stages.1 The mast of abnormal cells that grow in or around the brain is called a brain tumour.
It poses a risk to the healthy brain by either destroying or invading normal brain tissue. The tumour in the brain is emerged due to the existence of a glial cell known as GLIOMAS. Those cells are classified and graded from values 1 to 4. In this assigned grade, tumour belongs to grades 3 and 4 are stated as malignant or cancerous cell. The tumour belongs to grade 1 and 2 is stated as benign or non-cancerous cells.2 To identify brain tumour Magnetic Resonance Imaging (MRI) is utilized for the detection of modality to assess tumours in the brain. MRI assists the physician in the investigation of soft tissues in the human brain. MRI offers soft tissues with four different types such as T1-weighted (T1w), T1-weighted with contrast enhancement (T1wc), T2-weighted (T2w), and Fluid Attenuated Inversion Recovery (FLAIR).3 In this, healthy tissue is stated as T1-weighted (T1w).
Corresponding Author: M.Sumithra, Research scholar, Department of computer science and engineering, Sathyabama institute of science and technology, Chennai, India.
Both T2w and T1w are used for the detection of tumours which provides a bright tumour border. The FLAIR is involved in the isolation of the oedema region in the brain from CSF (cerebrospinal fluid). However, the identification of boundaries of the tumour is difficult due to homogeneity with different sequence intensities.4
Identification of brain tumour from MRI consist of different stages. Segmentation is termed to be a significant but tough step for the classification of medical imaging and its analysis.5 To segment MRI segmentation Convolutional Neural Networks (CNNs) are utilized with multi-modal factors. The CNN model consists of several functions such as extraction and classification of feature with a single model. However, existing CNN is subjected to complexity issues and limited accuracy.6 In recent years, Deep Convolutional Neural Network (DCNN) strategy is adopted for the extraction and classification of features. In, proposed a DCNN model was integrated with convolutional kernels for tumour segmentation in MRI images. In DCNN, a small kernel filter is adopted for cascade connection of convolutional layers with small kernel filters. 7 In developed an architecture with a parallel cascade connection of CNN. The cascaded network incorporates training with balanced classes and the second stage involved in the refinement of the last layer with several samples at each class.8
In developed an algorithm based on utilization of fuzzy c-means clustering through the utilization of membership function.9 Based on estimated membership function centres are clustered and simultaneously generated. Recently, in 10 presented a modified FCM algorithm for segmentation of MRI image. We developed a fuzzy segmentation for MRI images. The MRI image segmentation is based on the estimation of IIH with the characterization of Gaussian function. 9 A research conducted developed a multi-objective framework for 3D MRI image segmentation. The proposed approach incorporates a two-stage fuzzy multi-objective framework (2sFMoF). Also, in 13 constructed an FCM algorithm spatial information algorithm for segmentation noisy MRI images. Also, it contains local membership as an objective function for MRI image segmentation. 10,11 In developed a conditional spatial FCM(csFCM) for segmentation of MRI brain image. Similarly, in15 constructed a deep convolutional neural network for MRI image segmentation with the exploitation of convolutional kernels. However, this technique fails to provides an accurate classification of brain tumours. To overcome those limitations this research presented a DenseAlexNet model with a backpropagation classifier. evaluation metrics. 12,13
This paper developed a Dense AlexNet for tumour segmentation for brain tumour diagnosis. Also, the backpropagation scheme is applied for the classification of tumour regions in MRI images of the brain. 14
3D Dense Alex Net for MRI Segmentation
To achieve segmentation accuracy of 3D brain MRI images this paper uses 3D DenseAlexNet mode. The data for Multimodal Brain Tumour Segmentation is based on MICCAI 2012 conference. The developed Dense AlexNet model performance is based on the estimation of directions in a 3D image. The Dense AlexNet is involved in the extraction of features from brain images with the discriminative representation of MRI images through pooling layers. The next layer of Dense AlexNet generates high-level MRI image features for categorizing features in MRI images. In the next stage, the processed samples are masked for the identification of high-level features of MRI 3D images. With the object localization process in 3D Dense AlexNet, middle features are processed within the network. Through the incorporation of the backpropagation tumour region of the MRI image is segmented. To process input data MRI brain image is considered for pre-processing and segmentation of the tumour part. The feature of MRI brain images is extracted based on GLCM features with selected Region of Interest (ROI) for segmentation of the tumour part. The developed Dense AlexNet based on the adjusted pixel values tumour is segmented concerning position and area. Finally, classification is performed with Backpropagation for obtaining a resultant image with dataset images to identify it is benign or malignant. Initially, the proposed Dense AlexNet perform image pre-processing for removal of noise. To enhance the quality of the image certain features are examined to display image processing. The process involved in MRI pre-processing of MRI brain images is filtering of noise, pseudo-colouring, sharpening, and magnifying. The steps involved in improving image quality are image display, image analysis, and feature extraction. This paper uses median filtering for processing for the elimination of noises in the image. The applied median filter performs a linear operation for the elimination of salt and pepper noise. The process of median filtering is involved in the reduction of noise to preserve image edges. The proposed Dense AlexNet examines the image feature pixel, weight, depth, and colour before classification steps. Image segmentation is involved in the identification of object boundaries and location. Image explicit segmentation in involved in each pixel allocation based on assigned labels with similar label characteristics. Image segmentation outcome covers a complete image or performs image extraction. Every image pixels are based on characteristics of image property, texture, colour, and intensity. The Dense AlexNet performs classification of the image with backpropagation for MRI image training and testing. The proposed Dense AlexNet perform image classification using weighted estimation. The segmented images are trained and tested for image classification. The performance of the proposed Dense AlexNet is compared with image datasets. With image dataset classification tumour region of MRI images are segmented and classified as either normal or abnormal tumour images.
RESULTS
The performance of the proposed Dense AlexNet uses a Backpropagation classifier for segmentation and classification of tumour in MRI images. The proposed Dense AlexNet is implemented using MATLAB 2019 for analysis. To segment tumour in MRI image, Dense AlexNet is designed and obtained results are presented as follows. In figure 1 and 2, the input image considered for segmentation of the MRI brain image and figure 2 provides the pre-processed image for the input 3D MRI image.
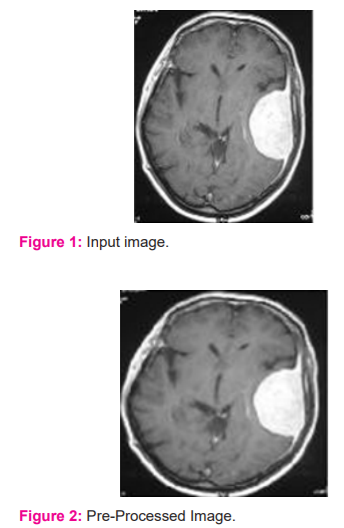
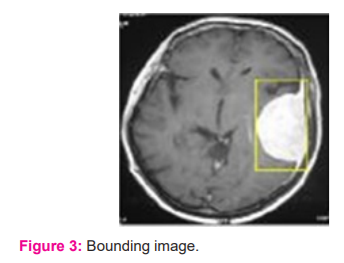
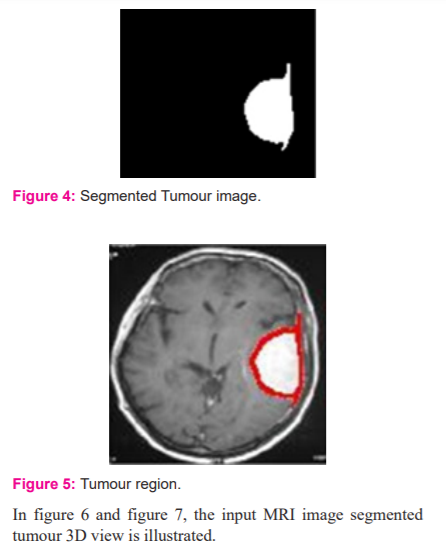
In figure 3, the bounding box for the pre-processed image is presented. The Dense AlexNet involved in the processing and quality enhancement of input 3D MRI images. To highlight the tumour within the input image bounding box is adopted. The bounding box estimates the image features and highlights the tumour region. In figure 4 segmented tumour region is presented for input 3D MRI brain image. In figure 5 detected tumour region of the proposed dense AlexNet is presented.
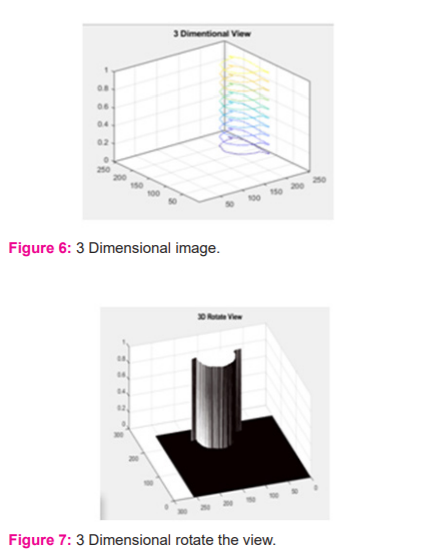
DISCUSSION
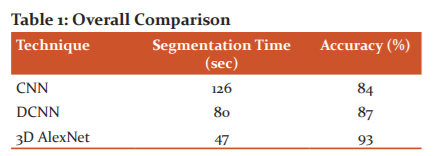
The developed 3D AlexNet model involved in the segmentation of 3D MRI brain images for segmentation. The developed 3D AlexNet model utilizes a backpropagation approach for segmentation of 3D MRI. Based on identified region proposed Dense AlexNet estimate the location of the tumour in the input MRI image and segment tumour part. The proposed Dense AlexNet significantly estimate the 3D view of the input image based on consideration of the input 3D MRI image of the brain for segmentation of tumour in input 3D MRI image. The comparative analysis of results expressed that the proposed 3D AlexNet model exhibits higher accuracy rather than other existing technique. In table 1, time and accuracy of proposed 3D AlexNet with existing Convolutional Neural Network (CNN) and Deep Convolutional Neural Network (DCNN). Table 1 shows the overall comparison.
In figure 8 comparative plot for segmentation time and accuracy is presented
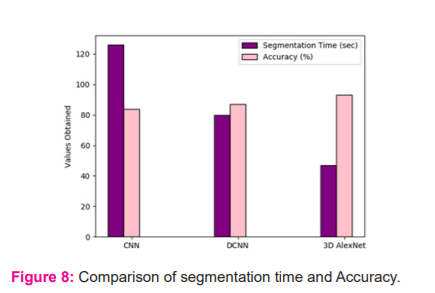
The simulation analysis expressed that the proposed 3D AlexNet model exhibits a higher accuracy value than the existing CNN and DCNN. Also, the proposed 3D AlexNet significantly reduces segmentation time compared with CNN and DCNN.
CONCLUSION
A brain tumour is caused due to the anomalous growth of tissues within the human brain, this leads to increase mortality. The proper diagnosis is required for reducing mortality rate, hence image processing techniques are evolved for effective diagnosis of brain tumours. Usually, the processing of MRI brain images is a complex task due to the complexity of the human brain structure. To improve the diagnosis process in MRI brain images, this paper proposed a Dense AlexNet for the segmentation of tumours. The proposed Dense AlexNet uses a Backpropagation classifier for tumour detection in the region. The comparative analysis of proposed 3D AlexNet architecture with conventional CNN and DCNN exhibited that improved accuracy with reduced segmentation time. The accuracy of the proposed 3D AlexNet is ~3% improved. Similarly, segmentation time is reduced on average by ~30 sec. In the future, the proposed segmentation scheme is processed with a sophisticated algorithm for detection, segmentation, and classification of brain tumour for medical applications.
ACKNOWLEDGMENT
The authors acknowledge the immense help received from the scholars whose articles are cited and included in references to this manuscript. The authors are also grateful to authors/editors/publishers of all those articles, journals, and books from which the literature for this article has been reviewed and discussed.
Conflict of Interest: Nil
Source of Funding: Nil
References:
-
Bondy M. L, Scheurer M. E, Malmer B, Barnholtz Sloan J. S, Davis F. G, Il'Yasova D and Sadetzki S. Brain tumosur epidemiology: consensus from the Brain Tumour Epidemiology Consortium. Cancer 2008;113: 1953-1968.
-
Havaei M, Davy A, Warde-Farley D, Biard A, Courville A, Bengio Y and Larochelle H. Brain tumour segmentation with deep neural networks. Med Ima Analy. 2017;35: 18-31.
-
Bauer S, Nolte L. P and Reyes M. Fully automatic segmentation of brain tumour images using support vector machine classification in combination with hierarchical conditional random field regularization. International Conference on Medical Image Computing and Computer-Assisted Intervention 2011; 354-361.
-
Prastawa M, Bullitt E, Ho S and Gerig G. A brain tumour segmentation framework based on outlier detection. Med Ima Analy. 2004;8(3):275-283.
-
Sumithra M and Malathi S. A survey on Medical Image Segmentation Methods with Different Modalitites. Int J Engg Res Techn. (IJERT) 2018; 6(2).
-
Sumithra M and Malathi S. A Survey of Brain Tumour Segmentation Methods with Different Image Modalitites. Int J Compt Sci Tren Techn (IJCST) 2017; 5(2).
-
Zhang C, Li P, Sun G, Guan Y, Xiao B and Cong J Optimizing fpga-based accelerator design for deep convolutional neural networks. Proceedings of the ACM/SIGDA international symposium on field-programmable gate arrays, 2015; 161-170.
-
Kamnitsas K, Ledig C, Newcombe V. F, Simpson J. P, Kane A. D, Menon D. K and Glocker B Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med Ima Analy. 2017;.36: 61-78.
-
Pal N. R, Pal K, Keller J. M and Bezdek J. C A possibilistic fuzzy c-means clustering algorithm. IEEE transactions on fuzzy systems. 2005;13(4):517-530.
-
Ji Z, Xia Y, Sun Q and Cao G Interval-valued possibilistic fuzzy C-means clustering algorithm. Fuzzy Set Syst 2014;138-156.
-
Mahata N, Kahali S, Adhikari S. K and Sing J. K Local contextual information and Gaussian function induced fuzzy clustering algorithm for brain MR image segmentation and intensity inhomogeneity estimation. Appl Soft Compt. 2018; 68:586-596.
-
Kahali S, Adhikari S. K and Sing J. K. A two-stage fuzzy multi-objective framework for segmentation of 3D MRI brain image data. Appl Soft Compt. 2017;60: 312-327.
-
Singh C and Bala A. A local Zernike moment-based unbiased nonlocal means fuzzy C-Means algorithm for segmentation of brain magnetic resonance images. Expert Syst Applic. 2019; 118:625-639.
-
Adhikari S. K, Sing J. K, Basu D. K and Nasipuri M. Conditional spatial fuzzy C-means clustering algorithm for segmentation of MRI images. Appl soft compt. 2015;34:758-769.
-
Pereira S, Oliveira A, Alves V and Silva C. A. On hierarchical brain tumour segmentation in MRI using fully convolutional neural networks: a preliminary study. IEEE 5th Portuguese meeting on bioengineering (ENBENG), 2017;1-4.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License