IJCRR - 13(11), June, 2021
Pages: 199-205
Date of Publication: 04-Jun-2021
Print Article
Download XML Download PDF
Molecular Assessment of Genetic Diversity Among Male, Female and Hermaphrodite Simarouba species Using Random Amplified Polymorphic Deoxyribose Nucleic Acid Markers
Author: Vaidya Gayatri, Naik GR
Category: Healthcare
Abstract:Introduction: The Simarouba species is considered one of the promising feedstock for biodiesel producers as the seeds have high oil content and are non edible. These species can be grown well even in wasteland. As Simarouba is polygamodioecious in nature and have a long juvenile period assessment of genetic variability in Simarouba germplasm is essential for exploitation of genetic resource for plant improvement programs. Objective: The present investigation aims to find the genetic diversity of male, female and andramonieocus Simarouba species using RAPD markers. Methods: Fifty Random amplified polymorphic DNA primers were used for genetic analysis of male, female and hermaphrodite Simarouba species. The binary data obtained from the RAPD banding profile were analysed using Wards Squared Euclidean distance and construct dendogram through UPGMA (Unweighted Pair Group Method with Arithmetic average statestica). Jac�card similarity coefficient was constructed using NTSYS ( Numerical Taxonomy System, PC version 2.02e programme to determine genetic relation among Simaroba individuals. Results: RAPD primers yielded 223 amplicons of which 176 (78.92 % ) were polymorphic. An average number of bands per polymorphic primer observed is 4.46. Jaccard Similarity Matrix between male, female and hermaphrodite Simarouba genotypes varied from 0.06 to 0.81. Among the various combinations, maximum similarity of 81 % was observed between Female and hermaphrodite individuals. While maximum dissimilarity of 63 % was found between female and male individuals. Dendrogram generated by UPGMA (Unweighted Pair Method with Arithmetic Mean ) grouped female and male plants in separate groups. Conclusion: Molecular markers employed in this investigation showed genetic diversity between crop genotypes. This study provides development of strategies for genetic analysis and crop improvement in Simarouba species.
Keywords: Crop improvement, Genetic diversity, Polymorphism, RAPD
Full Text:
Introduction
Simarouba glauca belongs to the family Simaroubaceae. It is a fast-growing, very important seed oil yielding multipurpose tree that grows even on marginal lands under water stress conditions.1 it also improves soil health. It is indigenous to the Amazon rainforest and other tropical areas in Mexico, Cuba, Haiti, Jamaica, and Central America; it was brought to India from Latin America in 1960. Simarouba species aid to control soil erosion and play a promising role in the conservation of soil and watersheds. The root system of the Simarouba species is well developed, which checks soil erosion, thus assisting soil microbial life and improving groundwater position. Plantation of these species facilitates wasteland reclamation, reduce greenhouse effect.2 These species break the wind.3 The oil cake being rich in nitrogen (8%), phosphorus (1.1%) and potash (1.2%) is good organic manure.2 The use of Simarouba press cake as a fertilizer has given good results on coffee, sugarcane, cotton and corn.4 The fruit pulp may be used in the production of vermicompost which is of excellent quality. Leaf litter makes good manure.5 Leaf litter (about 20kg/tree/yr.) makes good manure, improving soil fertility.6 Each well-grown tree yields 15-30 kg nutlets equivalent to 2.5 to 5.0 kg oil and about the same quantity of oil cake.5,7 The annual returns in kg/ha from a moderately well managed 10-year-old plantation of Simarouba is as follows. Oil:-1000-2000, Oil cake: - 1000-2000, Fruit pulp: - 8000-10000, Leaf litter:-10000-15000, Shell: - 4500-9000.6 The tree has high medicinal value used in pharmaceuticals, cosmetics and wood is used for furniture.8 Harnessing the full potential of non-traditional tree-borne oilseeds like Simarouba through systematic block plantations with elite material can carve out a niche that is noncompeting with current needs and support horizontal expansion of oilseeds, increase the green cover, reduce erosion, and generate employment in underdeveloped areas. The establishment of plantations of tree-borne oilseeds can solve the problems of conserving degraded lands, decrease the burden of shortages of fossil fuels and high energy import costs, and maintain a clean environment by reducing greenhouse gas emissions. This would require systematic efforts towards tree improvement program. An assessment of genetic variability in Simarouba germplasm is essential for the exploitation of genetic resource for plant improvement programs.
Morphological features, biochemical and cytological studies have not solved the problem of germplasm characterization determination in the economically important dioecious crops. Different attempts to find molecular differences between male and female plants such as immune chemistry, differences in protein and enzyme, mRNA and tRNA polymorphism or RNA hybridization kinetics have been made in several diocious species9 these studies deal with gene expression and could just show the differential expression of shared genes between male and female plants. This has led to the advancement of strategies. Plant biotechnology is enriched with multidisciplinary approaches of DNA/RNA fingerprinting techniques. In recent years, this approach has been applied to develop gender-specific DNA and RNA markers with greater reliability and accuracy and proved useful for marker-assisted selection of male and female and hermaphrodite plants. The stage of plant growth and the environment, in which the plant grows, do not influence the differences in DNA sequence. Therefore, molecular markers are currently being used to study of genetic diversity and identity of crop genotypes.10
Random amplified polymorphic DNA (RAPD) which is a polymerase chain reaction-based DNA fingerprinting technique.11,12 Differential PCR amplification of random genomic fragments using random decamer oligonucleotide primers are the basis of RAPD. DNA polymorphisms detected result from rearrangements or deletion at or between oligonucleotide primer binding sites in the genome. RAPD technique does not require sequence information of the genome that is being analyzed so it can be applied across species as universal primers.13 It has numerous applications such as identification of different cultivars genotypes, phylogenetics studies, diversity studies, selection of specific trait of interest, linkage mapping, etc.13-17 No prior knowledge of sequence information, cost efficiency, simple assaying procedure, less sophisticated equipment and no need for cloning and radioactive probes have made RAPD technique a valuable tool. In the present study, RAPD markers were used for molecular characterization and genetic variability of female, male, and hermaphrodite Simarouba plants.
Materials and methods
Morphological identification of sex of Simarouba plant by flower study
Immature leaf samples of both female (P1and P2) and hermaphrodite (P3 ad P4) plants were collected from University of Agricultural Science, Gandhi Krishi Vignan Kendra Bangalore. India. Male (P5) plant sample was collected from the plantation of Simarouba from University of Agricultural Sciences, Dharwad. India, the samples were stored at -80oC. Leaf sample was picked from male, female, and hermaphrodite plants after complete observation of flower types that were used for DNA extraction. The androecium of male and bisexual flowers have ten stamens in two whorls, where as in female flowers, the androecium is represented by ten staminodes which are sterile stamens with poorly developed anthers not producing pollen grains.
RAPD study
Isolation of genomic DNA from Simarouba species
Total genomic DNA was isolated from leaf tissues with the minor modifications in the CTAB method.18 About 0.3 g of leaf tissue was ground to a fine powder in liquid nitrogen and mixed with 700 µl of CTAB (cetyltrimethylammonium bromide) extraction buffer (100 mM Tris–HCl pH 8, 1.4 M NaCl, 20 mM EDTA pH 8, 2% CTAB, 1% b-mercaptoethanol, 1% PVP). This mixture was first incubated at 650C for 30 min, and then an equal volume of a phenol: chloroform: isoamyl alcohol (25:24:1) mixture was added, followed by centrifugation at 4,000 rpm for 30 minutes at 40C. The aqueous phase was decanted and transferred to a new microtube to reduce impurity between the two phases. The extraction steps were repeated using the same phenol: chloroform: isoamyl alcohol (25:24:1) mixture. The last aqueous phase was mixed with two-thirds volume of isopropanol and stored at -200C for at least 2 hours to precipitate the DNA, then centrifuged at 4,000 rpm for 15 minutes. The nucleic acid precipitate was washed with 70% ethanol, air-dried, and suspended in 50 µl of TE buffer.
Purification of DNA
The extracted DNA was treated with RNase (1mg/ml) (Qiagen, USA) incubated at 37oC for 1 hour. This was followed by Proteinase K (20mg/ml) treatment incubated at 37 oC for 30 minutes and sodium chloride to remove RNA, proteins, polysaccharides, and phenols, respectively.
Quantification of extracted DNA
The purity and quantity of the isolated DNA was determined by NanoDrop Spectrophotometer (ND 1000; Thermo Scientific, Wilmington, DE). The spectral ratio (A260 nm /A280 nm) was measured. The absorbance 1.0 at 260nm and 280nm determines the concentration of the nucleic acid (DNA) of 50 µg/ml in the sample. The ratio between the readings at 260 nm and 280 nm provides an estimate of the purity of nucleic acid. DNA preparation is pure when OD260/OD280 value is 1.8. If there is contamination with protein or phenol the OD260/OD280 will be slightly less than 1.8 and if more than 1.8 then it is contaminated with RNA
Total quantity of the DNA (µg/ml) = OD260 x 50 x dilution factor / 1000
Agarose gel electrophoresis
Agarose gel electrophoresis performed to check the quality of the extracted DNA samples from the plant samples. Agarose 0.8% (w/v) was prepared in 1x TAE buffer pH-8.0 (for 50x TAE, 242g Tris base, 37.2g ETDA, pH is adjusted with glacial acetic acid).The mixture is heated till agarose is completely dissolved and forms a clear solution. The molten agarose is poured in the gel mould which was set with the comb when cooled down at about 55oC. The set up was left for an hour at room temperature for solidification. The gel slab is placed in an electrophoresis unit, 1 x TAE buffer was poured to immerse the gel. DNA samples were prepared by mixing with 10µl of tracking dye Bromophenol Blue (50% Glycerol, 1Mm ETDA-pH-8.0, xylene cyanol and distilled water) and the samples were loaded with uncut λ DNA (Bangalore Genei, Bangalore, India) of known concentration in to the wells. Agarose gel was run for about two to three hours at a constant voltage of 5V/cm2 in electrophoresis tank. The gel was visualized under Gel Documentation System. The high intensity bands were considered as good quality DNA samples and smear were of poor quality.
Selection of RAPD Primers
Fifty RAPD primers were obtained from Eurofins MWG Biotech, Bangalore India. These primers were screened with extracted DNA of male, female and andromonecious Simarouba plants. RAPD data analysis was done using only the amplifying primers.
PCR amplification
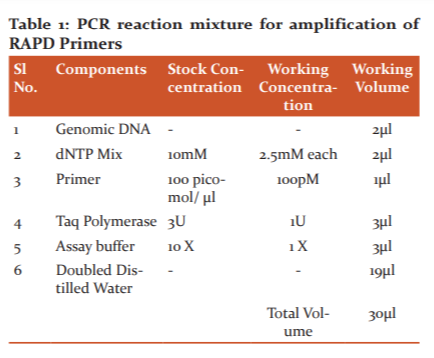
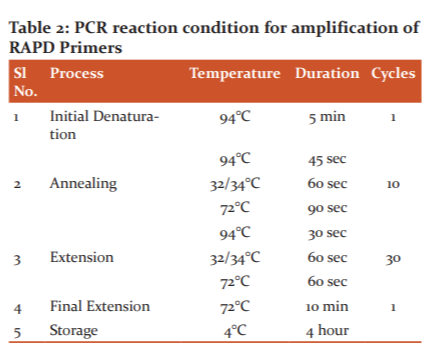
Fifty 10-base primers (Eurofins MWG Biotech, Bangalore India) were used for polymerase chain reaction (PCR) to screen known sex to ascertain their potential of clear amplification in polymorphism and the reproducibility. The RAPD-PCR reactions were performed in 30 μl volumes in 100 μl PCR tubes (Tarson Pvt Ltd, India) (Table 1). The reaction mixture contained 30 ng of template DNA, 1 × amplification buffer (10 mM of Tris-HCl- pH 8, 50 mM of KCl, 1.8 mM of MgCl2 and 0.01 mg/ml gelatine), 2.5mM each of dCTP, dGTP, dATP, and dTTP, 5pM primers and 1U Taq DNA polymerase (Bangalore Genei, Pvt. Ltd., India). The RAPD-PCR analysis was performed in a 30 µl volume in gradient Thermo Cycler. The reactions were performed in a Master Cycler Gradient 5331 (Eppendorf version 2.30. 31-09, Germany). The reaction had an initial denaturation step at 940C for 5 min, followed by 35 cycles of 94 oC for 1 min, 37oC for 1 min, 72oC for 2 min. The final extension step was at 72 oC for 10 min. The reactions were cooled and held at 4oC (Table 2).
The RAPD-PCR products were separated on 1.5% (w/v) agarose (Sigma-Aldrich, USA) gel at 5 V/cm in 1 × TBE (89 mM Tris-HCl, 89 mM boric acid and 2 mM EDTA, pH 8.0) buffer. The agarose gels were stained with 0.5 μg/ml ethidium bromide, visualized under UV light and photographed on a digital gel-documentation system (SYNGENE). The molecular weights of the RAPD amplicons were estimated with a 100 bp DNA ladder.
Data Analysis
Scoring of RAPD banding profile
The RAPD allele size was determined based on the location of bands with respect to the ladder. The total no of allele was recorded for each RAPD marker in all the accessions by giving allelic number 1, 2, 3, 4, 5, etc. Distinct visible and reproducible bands were scored as 1 for the presence and 0 for absence in DNA profile photographs. Differings band intensities were not considered to avoid errors introduced by competition among priming sites during the initial rounds of RAPD-PCR analysis.19
Genetic similarity and cluster analysis
The binary data obtained from the RAPD banding profile were analysed using Wards Squared Euclidean distance, and was used to construct dendogram through UPGMA (Unweighted Pair Group Method with Arithmetic average statestica). Jaccard similarity coefficient was constructed using NTSYS (Numerical Taxonomy System, PC version 2.02e programme to determine genetic relation among Simaroba individuals.
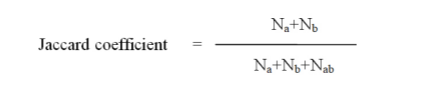
Where Na is the number of fragments present in the pattern a but not b.
Nb is the number of fragments present in the pattern b but not a.
Nab is the number of fragments commen for pattern a and b.
Results
Morphological study of flowers of Simarouba to identify the gender
Simarouba is a polygamodioecious with three types of plants, pistillate, staminate and andromonoecious. The inflorescence is compound panicle with main axis showing racemose pattern and ultimate branches having dichasial / monochasial cymes. The time taken from bud initiation to anthesis is about 15 to 20 days. The male inflorescence has longer peduncle than the female. The length of panicle is 12-40 cm in pistillate whereas 45-55 cm in andromonoecious and staminate inflorescences. The bisexual flowers are the largest with 10-15 mm in diameter, followed by male inflorescence with 8—12 mm diameter, and the female flowers are the smallest with 5-7 mm diameter. Female flowers were characterized by sterile stamens and anthers, which did not showed pollen grains whereas male and bisexual flowers showed active stemens with pollen grains production.
Isolation of Genomic DNA
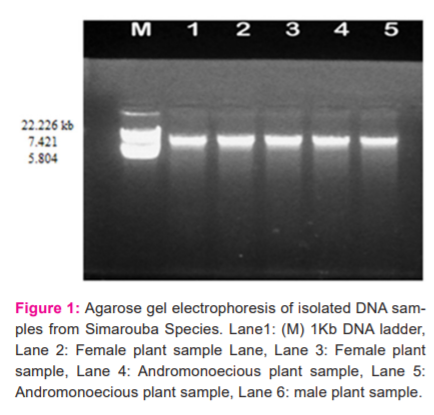
Genomic DNA was isolated by the CTAB method, purified by using RNase and proteinase treatment. it was analysed by agarose gel electrophoresis. The λDNA EcoRI digest (Merk, Bangalore) marker was used to determine the amplified DNA's molecular weight. The high quality DNA suitable for RAPD analysis was observed, intact discrete bands of high molecular weight were seen in the agarose gel ( Figure 1).
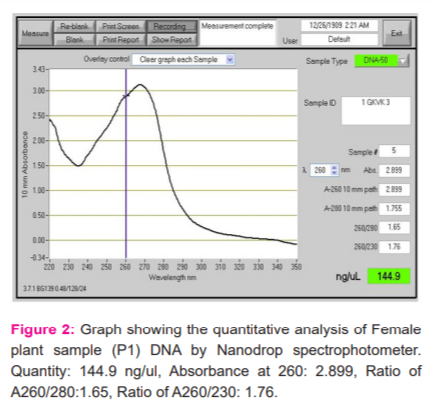
Quantification of Isolated Genomic DNA
The quantification of DNA was determined by Nanodrop spectrophotometer. The spectral ratio (A260/A280)of the DNA isolated from Simarouba individuals was found to be in the range of 1.8 to 2.1, which is an indication of low polysaccharides and high quality (Figures 2-6).
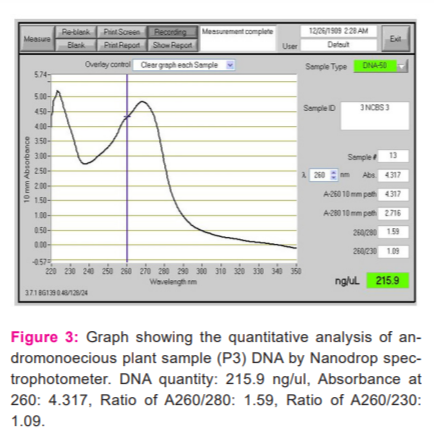
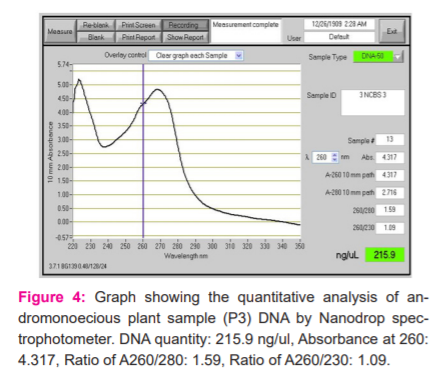
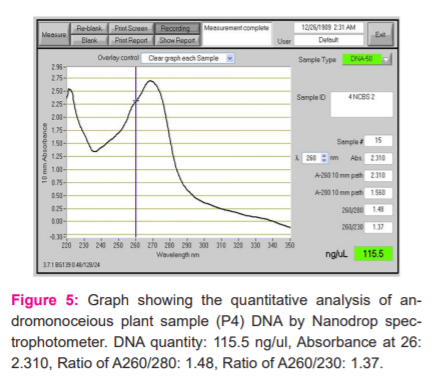
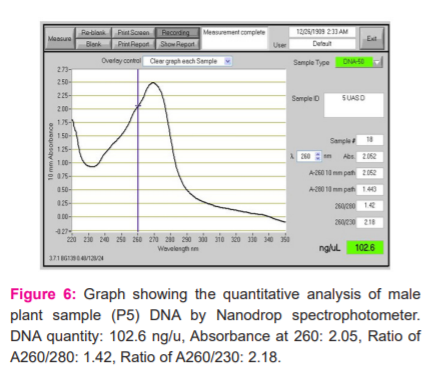
RAPD-PCR Analysis of Simarouba species.
RAPD-PCR has widely been used for population genetic studies as well for various amplification in the several plants. Attempts are made to screen for the primer, which would produce reproducible banding patterns for the genetic analysis of female, male, and hermaphrodite Simarouba plants. A set of 50 decamer primers were used to amplify the genomic DNA of male, female, and hermaphrodite individuals, of which 38 primers showed reproducible results.
Statistical Analysis
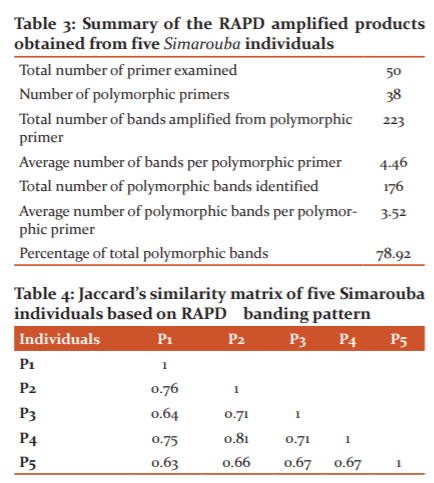
Total of 50 primers were screened to amplify the genomic DNA of Simarouba plants. The total number of bands amplified from polymorphic primers were 223, the percentage of total polymorphic bands is 78.92% and a number of amplified bands per primer are 4 (Table 3).The binary data of RAPD-PCR amplification was analysed using Wards squared Euclidean distance. Dendrogram analysis was done using gel documentation system. Phylogenetic variation was determined by converting RAPD data into Jaccard's Similarity Coefficient and analyzed by unweighted pair method with arithmetic mean (UPGMA) cluster analysis to produce phylogenetic tree. The similarity matrix (Table 4) reveals that the cultivars fall in the range of 0.63 to 0.81. Among the various combinations, maximum similarity of 81% was observed between P2 and P4 (Female and Andromonecious) cultivars, while maximum dissimilarity of 63% was found between P1(Female) and P5 (Male) cultivars.


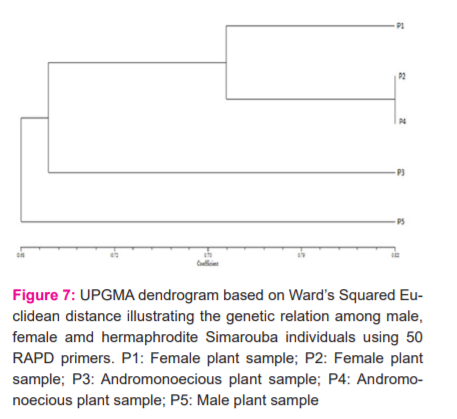
Dendrogram generated (Fig.7) based on RAPD data, divided Simarouba cultivars into two distinct clusters. Cluster I comprises of four plants, namely P1, P2, P3 and P4. Cluster II comprises of one plant namly P5. Cluster I is subdivided in to three sub-clusters (I, II, III), sub cluster I comprises of single cultivar that is P1, sub-cluster II comprises of two closely related cultivars those are P2 and P4 and sub-cluster III consist of single cultivar that is P3. The position of P1(female) and P5(male) are at extreme end on dendrogram indicating them as most divergent species. Cluster analyses based on Wards squared Euclidean distance shows combining results from fifty primers. On the linkage distance skill, it is evident that female and andrimoniceous plants are closely linked whereas male and female are divergent.
Discussion
In the present study, Random amplified polymorphic DNA primers were tested on diecious and hermaphrodite plants of Simarouba glauca. The Jaccard's similarity co-efficient constructed by binary data of the RAPD-PCR amplification was ranged between 0.63 to 0.81 among Simarouba individuals. The lowest coefficient (0.63 ) was found between male and female plant and the highest similarity coefficient (0.81) was found between female and andromonoecious plants. This reveals the least similarity between male and female plants. Dendrogram was constructed to determine the genetic relationship among the Simarouba individuals. Cluster analysis based on Squared Euclidean distance is represented by the results obtained from oligo primers. On linkage distant scale, it is evident that female and andromonoecious are closely linked. Male is distinctly linked to both female and andromonoecious. The individual of each sex does not form separate cluster; instead, they are mixed, indicating that they are evolution. The most extreme type of gender determination system is found where highly specialized gender chromosomes are found, which usually promotes dioecy in plants.20 The evolution of dioecy from the hermaphrodite species is considered unlikely, since the occurrence and establishment of two independent mutants, one for female and other for male sterility must occur simultaneously, and the mutant genes or multiple loci must be tightly linked so that generation of the hermaphrodites does not occur by recombination.21 As dioecism has raised in different families and genera of plants22 the development of molecular strategies of dioecious taxa has been a priority in breeding programmes for their greater economic potentials Marker-based differences in the genetic relationships between Simarouba genotypes showed that cluster tree analysis has grouped of male and female plants genotypes in separate groups
Conclusion
The use of molecular markers to distinguish the genders has been employed when the genetic mechanism of gender determination is not available. Molecular marker-based technology has proved a reliable strategy for detecting gender-associated markers in dioecious and bisexual plants. The RAPD marker technique is the cheapest, user friendly and reliable tool used for efficient fingerprinting of many plants. The oligo primers used in the present study were found consistent and reproducible. RAPD marker technique proved to be useful in the present investigation. Studies on marker technology regarding dioecy have rendered a better understanding of dimorphism's development and evolutionary process.
Acknowledgments: Author gratefully acknowledges Department of Biotechnology Gulbarga University Gulbarga for providing the work felicity and Dr Shyam Sundar Joshi retired professor from Department of Botany University of Agricultural sciences. Bangalore. India to provide relative information and Professor S. J. Patil from Department of farm forestry University of Agricultural sciences. Dharwar. India to help for procuring the study materials.
Conflict of interest: Authors declares that there is no conflict of interest.
Source of funding: There is no any source of funding or the present study
Authors contribution: Prof G R Naik have framed the study and cross checked the findings, and Dr Gayatri Vaidya has performed the experiments and analyzed the data, and prepared the manuscript.
References:
1. Rout GR. Das P. In vitro micropropagation of mature S. glauca LINN. an oil yielding tree. Bangladesh J Bot. 1995;24:137-14.
2. Patil MS, Gaikwad DK. A Critical Review on Medicinally Important Oil Yielding Plant
Laxmitaru (Simarouba glauca DC.). J Pharm Sci Res. 2011;3(4):1195-1213.
3. Raina SN, Rani V, Kojima T, Ogihara Y, Singh KP, Devarumath RM. RM RAPD and ISSR fingerprints as useful genetic markers for analysis of genetic diversity, varietal identification, and phylogenetic relationships in peanut (Arachis hypogaea) cultivars and wild species. Genome 2001;44:763–772.
4. de Sola F. Notes on the aceituno tree (Simarouba glauca DC.) and its adaptation as a vegetable oil crop. Ceiba. 1956;4:351–358.
5. Govindaraju K, Darukeshwara J, Srivastava AK. Studies on protein characteristics and toxic constituents of Simarouba glauca oil seed meal. Food Chem Toxicol. 2009;47:1327-1332.
6. Joshi S, Joshi S. OIL TREE- Laxmitaru glauca. University of Agricultural Sciences, Bangalore and Indian Council of Agricultural Research, New Delhi, India. 2002:86.
7. Kaul S. Biodiesel: a clean and sustainable fuel for future. In: All India seminars on national policy on – non-edible oils as biofuels. IISC Bangalore. 2003.
8. Gilman EF, Watson DG. Simarouba glauca Paradise Tree, Institute of food and Agricultural 210 Sciences, University of Florida, Gainesville, FL 32611, Fact Sheet. 1994;211 ST-590.
9. Durand R, Durand B. Sexual determination, and sexual differentiation. Crit Rev Plant Sci. 1990;9:67–77.
10. Smith S, Helentjaris T. In Genome mapping in plants (AH Paterson, Editor), Academic Press, RG Landes Company. Texas. 1996; 95.
11. Welsh J, McClelland M. Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res. 1990;18:7213–7218.
12. Williams JGK, Rubelik AR, Livak KJ, Rafalski A, Tingey SV. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 1990;18:6531–6535.
13. Williams MNV, Pande N, Nair S, Mohan M, Bennett J. Restriction fragment polymorphism analysis of polymerase chain reaction product amplified from mapped loci of rice genomic DNA. Theor Appl Genet. 1991; 82:489–498.
14. Fernandez ME, Figueiras AM, Benito C. The use of ISSR and RAPD markers for detecting DNA polymorphism, genotype identification and genetic diversity among barley cultivars with known origin. Theor Appl Genet. 2002;104:845–851.
15. Iruela M, Rubio J, Cubero JI, Gil J, Millan T. Phylogenetic analysis in the genus Cicer and cultivated chickpea using RAPD and ISSR markers. Theor Appl Genet. 2002; 104:643–645.
16. Wong CL, Gan SY, Phang SM. Morphological and molecular characterization, and differentiation of Sargassum baccularia and S. polycystum (Phaeophyta). J Appl Phycol. 2004;16:439–445.
17. Yang X, Quiros C. Identification, and classification of celery cultivars with RAPD markers. Theor Appl Genet. 1993;86:205–212.
18. Samantaray S, Geetha KA, Hidayath KP, Maiti S. Identification of RAPD markers linked to sex determination in guggal [Commiphora wightii (Arnott.)] Bhandari. Plant Biotechnol Rep. 2010;4:95–99.
19. Bachmann K. Nuclear DNA markers in plant biosystematic research. Opera Botanica. 1997;132:137–148.
20. Ming R, Wang J, Moore PH, Paterson AH. Sex chromosomes in flowering plants. Am J Bot. 2007;94:141–156.
21. Ainsworth C. Boys and girls come out to play: the molecular biology of dioeciously plants. Ann Bot. 2000; 86:211–221.
22. Westergaard M. The mechanism of sex determination in dioecious flowering plants. Adv Genet. 1958;9:217–281.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License