IJCRR - 4(8), April, 2012
Pages: 55-62
Date of Publication: 25-Apr-2012
Print Article
Download XML Download PDF
COMPARATIVE MICROBIOLOGIC ANALYSIS OF SUBGINGIVAL PLAQUE SAMPLES IN TYPE II DIABETIC AND NON - DIABETIC PATIENTS WITH CHRONIC PERIODONTITIS BY POLYMERASE CHAIN REACTION
Author: Mythireyi D, M G Krishnababa, Kalaivani
Category: Healthcare
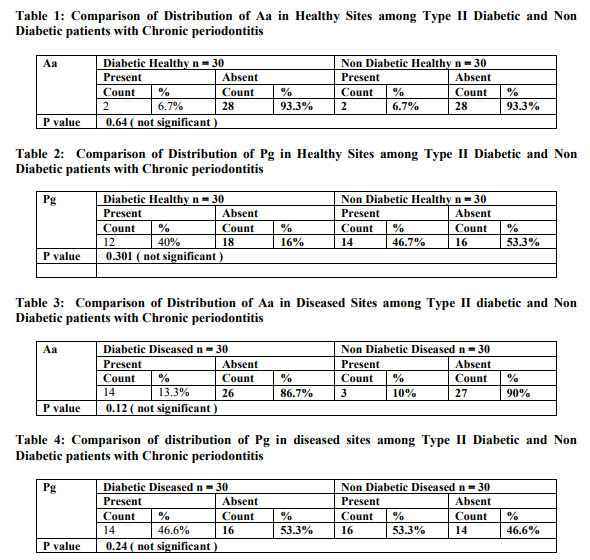
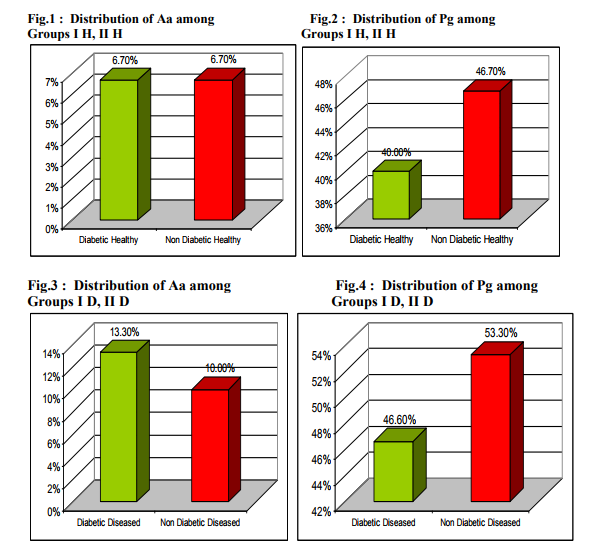
Abstract:Background: Although Immuno inflammatory relationship between periodontal diseases and diabetes mellitus is acknowledged, the difference in putative periodontal microorganisms between diabetic and non diabetic individuals is not well established. Aim: To compare the prevalence of two putative periodontal pathogens namely Aggregatibacter actinomycetemcomitans and Porphyromonas gingivalis in Type II Diabetic and Non Diabetic patients with chronic periodontitis by Polymerase chain reaction Materials and Method: Sixty subjects were selected from the Department of Periodontics, Tamilnadu Government Dental College and Hospital, Chennai \? 03. 30 Type II diabetic patients with chronic periodontitis were categorized as Group I and 30 Non diabetic patients with chronic periodontitis were categorized as Group II based on American Dental Association classification 1997 and American Academy of Periodontology classification 1999. Two sites- 1 healthy site and 1 diseased site were chosen in each patient, Group I H, II H \? healthy site samples and Group I D, II D- diseased sites samples. Subgingival plaque was collected, DNA isolation was done & the presence of A.actinomycetemcomitans & P.gingivalis DNA was determined by PCR. The PCR products were sequenced and confirmed. The data was statistically analysed. Results: A.actinomycetemcomitans was detected in 6.7 %, 6.7%, 13.3%, 10% in Groups I H, II H, I D, II D respectively. P.gingivalis was detected in 40%, 46.7%, 46.6%, 53.3% in Groups I H, II H, I D, II D respectively. When comparisons were made between Groups I H & II H and Groups I D & II D for the two organisms, no statistically significant difference was obtained Conclusion: The present study shows no statistically significant difference in the prevalence of A.actinomycetemcomitans and P.gingivalis in Type II Diabetic and Non Diabetic patients with chronic periodontitis.
Keywords: Aggregatibacter actinomycetemcomitans, Porphorymonas gingivalis, Diabetes, Periodontitis, Polymerase chain reaction
Full Text:
INTRODUCTION
Chronic inflammatory periodontal disease (periodontitis) is primarily an anaerobic Gram negative oral infection that leads to gingival inflammation, destruction of periodontal tissues, loss of alveolar bone and eventual exfoliation of teeth in severe cases 11. Certain organisms within the microbial flora of dental plaque are the major etiological agents of periodontitis. Traditional thinking / paradigms have maintained that periodontitis is an oral disease
and that the tissue destructive response remains localized within the periodontium. Whereas studies by Cohen DW et al 19704 , Mattila KJ et al 19898 have indicated that periodontitis may produce a number of alterations in systemic health. Diabetes mellitus is a metabolic disorder characterized by altered glucose tolerance or impaired lipid and carbohydrate metabolism 1 . It has been suggested that a positive correlation exists between diabetes and periodontal destruction based on the fact that loss of periodontal attachment occurs more frequently and more extensively in moderately and poorly controlled diabetic patients than those under good control. Diabetes mellitus influences prevalence and severity of periodontal disease. Although host immune inflammatory response plays an important role, it is the microflora that‘s proved to be the etiological agent in periodontitis
AIM
The aim of the present study was to compare the prevalence of two putative periodontal pathogens namely Aggregatibacter actinomycetemcomitans and Porphyromonas gingivalis in Type II Diabetic and Non Diabetic patients with Chronic periodontitis by Polymerase chain reaction.
MATERIALS AND METHOD
SUBJECT SELECTION
Sixty subjects were screened and selected from the out patient Department of Periodontics, Tamilnadu Government Dental College and Hospital, Chennai – 600 003. 30 Type II diabetic patients with chronic periodontitis were categorized as Group I (Study group) and 30 Non diabetic patients with chronic periodontitis patients were categorized as Group II (Control group). Within each group sub categorization was done as follows: Group I H - 30 Healthy sites of Type II Diabetic patients with chronic periodontitis Group I D - 30 Diseased sites of Type II Diabetic patients with chronic periodontitis Group II H - 30 Healthy sites of Non Diabetic patients with chronic periodontitis Group II D - 30 Diseased sites of Non Diabetic patients with chronic periodontitis The criteria for selection of patients with Type 2 Diabetes was based on American Diabetes Association classification (1997)1 and for Chronic periodontitis and Chronic periodontitis modified by Diabetes, the American Academy of Periodontology classification (1999)2 was utilized.
INCLUSION CRITERIA
Age 30 – 60 yrs, Either sex, At least 3 sites with PPD 7 mm, CAL >1mm, At least 2 sites with PPD ≤ 3mm, Type 2 Diabetic patients (Group I), Systemically healthy individuals (Group II)
EXCLUSION CRITERIA
Patients with systemic disease other than Type 2 diabetes(Group I), Patients with systemic disease (Group II), Antibiotic therapy for the past 6 months, Smokers, Periodontal therapy for the past 1 year
Following Institutional Ethical Committee Approval, selection of subjects was done. Informed consent was obtained and a thorough medical and dental history was taken. Intra-oral examination was done using mouth mirror and William‘s periodontal probe. Periodontal evaluation was done by measuring the Plaque Index, the Gingival Bleeding Index, Probing Pocket depth( PPD) and Clinical Attachment level(CAL).
Collection of subgingival plaque sample and polymerase chain reaction
Two plaque samples were taken from the most diseased site and a healthy site from each patient with individual sterile Gracey curettes. The plaque was dislodged into a vial containing 200 l of sterile lysis solution (10mm tris, 1.0 mm EDTA , 1.0% Tris X – 100, pH 78), sealed and stored at -20oC. Subgingival plaque samples was boiled for 10min, cooled to room temperature, centrifuged at 10,000 rpm for 3 min and the supernatant was stored at -20ºC till assay. 10µl of the supernatant was directly used as template in PCR.
Primers utilized in this study
Aggregatibacter actinomycetemcomitans (Aa)
Forward primer : 5‘ CAGCAAGCTGCACAGTTGCAAA – 3‘ Reverse primer : 5‘ CATTAGTTAATGCCGGGCCG TCT – 3‘
(Kraig E et al / Infect Immun 1900, 58 : 920 – 929)
Porphyromonas gingivalis (Pg)
Forward primer : 5‘ ATAATGGAGAACGCAGG AA -3 Reverse primer : 5‘ - TCTTGCCAACCAGTTCCA TTGC – 3‘
(Dickinson et al / J Bacteriol 1998 ; 170 ; 1658 – 1665)
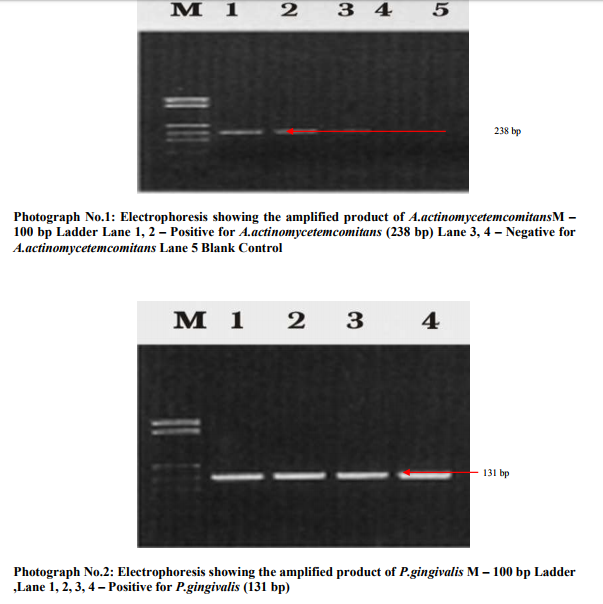
10 µl of the PCR master mixture was pipette into micro centrifuge tubes. 1.0 l of forward primer, 1.0 l of reverse primer, 3.0 l of template DNA and 5.0 l of nuclease free water was added and mixed thoroughly. The micro centrifuge tubes were placed in thermocycler and cycling conditions were set. PCR was performed for 35 cycles of Denaturation at 95oC for 1 min, Primer Annealing at 55oC for 30 sec, Primary extension at 72oC for 1 min, Final extension was 72oC for 10 min. The PCR product was detected by 2% Agarose gel electrophoresis. After the completion of the electrophoresis, gel was taken to the transilluminator and observed under UV-light. (Biorad gel documentation) The PCR product of 238 bp A.actinomycetemcomitans and131 bp P.gingivalis were given to MWG –
BIOTECH, GERMANY for sequencing the PCR products by automated DNA sequencer. The data was collected and statistically analyzed.
STATISTICAL ANALYSIS
Pearson‘s chi square test was used to calculate the overall p value The statistical package SPSS V18 (Statistical Package for Social Science, Version 18) was used for statistical analysis. In the present study, < 0.05 was considered as the level of significance.
RESULTS
In the present study A.actinomycetemcomintans was detected in 6.7% Type II diabetic patients with chronic periodontits -healthy sites (2 out of 30 healthy sites), 13.3% Type II diabetic patients with chronic periodontits -diseased sites(4 out of 30 diseased sites), 6.7% Non diabetic patients with chronic periodontits - healthy sites (2 out of 30 healthy sites) and10.0% Non diabetic patients with chronic periodontits -diseased sites(3 out of 30 diseased sites). In the present study P.gingivalis was detected in 40% Type II diabetic patients with chronic periodontits - healthy sites (12 out of 30 healthy sites, 46.6% Type II diabetic patients with chronic periodontits -diseased sites( 14 out of 30 diseased sites), 46.7% Non diabetic patients with chronic periodontits - healthy sites(14 out of 30 healthy sites) and 53.3% Non diabetic patients with chronic periodontits -diseased sites (16 out of 30 diseased sites DISCUSSION The clinical parameters used in this study were Gingival Bleeding Index, Plaque Index, Probing Pocket Depth and Clinical Attachment level similar to the study by Yuan K et al 200113. Earlier studies employed curettes or paper points for subgingival plaque collection. Sampling by paper point is less invasive than by curette but may result in an underestimation of tightly adherent bacteria in subgingival sites5, 12.Hence in this study curettes were used for sample collection. Although various diagnostic techniques are available to analyse the microbial population of subgingival plaque, PCR technique was opted as it can detect organisms of less than 100 cells3 . In the present study, PCR procedure followed by Yuan et al 200113 was used. Mandel et al 19907 detected 7% A.actinomycetemcomitans, 13% P. gingivalis in diseased sites of NIDDM patients by culture. Zambon et al 198813 employed culture and ELISA assays and detected P.gingivalis in 75% NIDDM subjects. In the present study, 13.3% and 46.6% diseased sites in Type II Diabetic patients with chronic periodontitis and 10.0% and 53.3% of diseased sites in Non Diabetic patients with chronic periodontitis were positive for A.actinomycetemcomitans and P.gingivalis respectively. Our microbiological data revealed higher detection rates when compared with the results of others (except Zambon et al). This may be due to the higher sensitivity of PCR, PCR is more sensitive than culture, immunofluroesence and DNA probes for which sensitivities are 2 x 102 , 2 x 105 , 2 x 104 , 2 x 103 respectively6,14. In a PCR study by Yuan et al 200113, 6.7% and 64.8% of diseased sites in Type II diabetic subjects and 5.7% and 66.7% of diseased sites in Non diabetic patients were positive for A.actinomycetemcomitans and P.gingivalis respectively. The discrepancy in results, may be because a large sample size of 150 patients was chosen by Yuan et al 200113 when compared to the small sample of 30 patients chosen in this study. Also, different authors have analyzed the microorganism distribution in different population and races. When comparing the prevalence rates of A.actinomycetemcomitans and P.gingivalis, the results in our study showed a lower detection rate for A.actinomycetemcomitans which is in concurrent to the study by Yuan K et al 200113 . This may be explained by the conclusion drawn from the studies by Rhodenburg JP et al 19909 , Slots J et al 199010 that the prevalence of A.actinomycetemcomitans was age related and decreased with increasing age. A.actinomycetemcomintans is more responsible for aggressive periodontitis whereas P.gingivalis contributes to chronic periodontitis. Since our subjects were all adults beyond 30 years of age, it is speculated that the contribution of A.actinomycetemcomitans to the periodontitis we examined was minimal. There was no statistically significant difference in the detection rates of the 2 tested microorganisms The plausible reasons are 1) There was no difference in the contribution of the microbiological pathogens in patients with Type II diabetic patients with chronic periodontitis and in Non diabetic patients with chronic periodontitis. 2) The PCR assay is limited in the ability to differentiate large or small amounts of the same pathogen. 3) Other microflora rather than our targeted microorganism such as P.intermedia, C.rectus and Capnocytophaga species (since they are also regarded as important pathogen in periodontitis of NIDDM patients13 )could be an etiological agent in Type II diabetic patients with chronic periodontitis and Non diabetic patients with chronic periodontitis.
CONCLUSION
The search for the etiologic agent of periodontal diseases has been in progress for over a century. In this study it was found that the composition periodontal microflora in periodontal disease sites of Type II diabetic patients with chronic periodontitis was similar to that found in non diabetic patients with chronic periodontitis.
However, the significantly higher detection rate of P.gingivalis in diseased sites further confirms the possible pathogenic role of this bacteria for both groups studied. A.actinomycetemcomintans may not be a causative pathogen in Type II diabetic patients with chronic periodontitis and non diabetic patients with chronic periodontitis. Also, PCR assay provides only a binary results i.e it detects the presence/absence of the microorganism and cannot differentiate positive result s quantitatively. Therefore the difference may exist in the quantitative composition of periodontal microorganism present in Type II diabetic patients with chronic periodontitis and non diabetic patients with chronic periodontitis. Hence further studies with a large sample size and diagnostic technique to quantitatively analyse the composition of microorganisms may be needed.
ACKNOWLEDGEMENT
Authors acknowledge the immense help received from the scholars whose articles are cited and included in references of this manuscript. The authors are also grateful to authors / editors / publishers of all those articles, journals and books from where the literature for this article has been reviewed and discussed.
References:
1. American Diabetes Association . Report of the Expert Committee on the Diagnosis and Classification of Diabetes mellitus Diabetes Care 2001: 24 (Suppl 1 ) S5 –S20
2. Armitage GC: Periodontal diagnosis and classification of periodontal diseases Periodontology 2000 :2004; 34;9 -21
3. Ashimoto A, I Bakker I Slots : Polymerase chain reaction detection of 8 putative periodontal pathogens in subgingival plaque of gingivitis and advanced periodontitis lesions. Oral Microbiol Immunol 1996; 11;266 -273
4. Cohen DW, Friedman LA, Shapiro J, Kyle GC, Franklin S Diabetes mellitus and periodontal disease Two year longitudinal observations Part I J Periodontol 1970 , 41; 709 -712
5. Hartroth B, Jeyfabrt I, Comado G Sampling of periodontal pathogens by paper points : Evaluation of basic parameters Oral Microbiol Immunol 1999; 14; 326 -330
6. Maiden MFJ, Tanner A, Mc Ardle S, Nagpauer K, Goodson JM Tetracycline fibre therapy monitored by DNA probe and culture methods J Periodont Res 1991 ;26;452-459
7. Mandell RL, Dirienzo T, Kent R, Joshipura K,Haber J Microbiology of healthy and diseased periodontitis sites in poorly controlled insulin dependant diabetes J Periodontol 1992;63; 274- 279
8. Mattila KJ, Nieminen MS, Valtonen W, Rasi VP, Kesaniemi YA, Syrjala SL Association between dental health and myocardial infarction BMJ 1989 ;298;779- 782
9. Rodenburg JP, Van Winkelhoff AJ, Winkel EG, Goene RJ, Abbas F, de Graff J Occurrence of Bacteroides gingivalis, Bacteroides intermedius and Actinobacillus actinomycetemcomitans in severe periodontitis in relation to age and treatment history J Clin Periodontol 1990; 17; 392-399
10. Slots J, Feik D, Rams JE Actinobacillus actinomycetemcomitans and Bacteroides intermedius in human periodontitis Age relationship and mutual association J Clin Periodontol 1990; 17;659 -662
11. Socransky SS and Haffajee AD The bacterial etiology of destructive periodontal disease current concepts J Periodontol 1992;63;4;322-331
12. TannerAC, Goodson JM Sampling of microorganisms associated with periodontal disease Oral Microbiol Immunol 1986; 1; 15 -22
13. Yuan K, Chang CJ, Hsu pc, Sun HS, Tseng CC, Wang Jr Detection of putative periodontal pathogens in non insulin dependant diabetes mellitus and diabetes mellitus by Polymerase chain reaction Periodont Res 2001; 36;18-24
14. Zappa u, Reinking – Zappa M ,Graf H, Comparison of serological and DNA probe analysis for detection of suspected periodontal pathogens in subgingival plaque samples Arch Oral Biol 1990; 35;161-164.

|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License