IJCRR - 4(22), November, 2012
Pages: 99-104
Date of Publication: 24-Nov-2012
Print Article
Download XML Download PDF
ANTIMICROBIAL SUSCEPTIBILITY PATTERN OF CLINICAL ISOLATES OF PSEUDOMONAS AERUGINOSA IN AN INDIAN TERTIARY CARE HOSPITAL
Author: A. Heraman Singh, Ranjan Basu
Category: Healthcare
Abstract:Objective: Resistance to antipseudomonal antibiotics is increasing worldwide. Area-wise studies on antimicrobial susceptibility patterns are essential to guide policy on the appropriate use of antibiotics. The present study was conducted to find out the current antimicrobial susceptibility pattern of P.aeruginosa isolates obtained from various clinical samples at our hospital. Methods and Material: The present study was conducted in a tertiary care hospital in Rajahmundry, Andhra Pradesh , India , during Jan 2011 \? Jun 2012. 107 clinical isolates of P.aeruginosa obtained from various clinical samples were studied. They were identified by Routine standard operative procedures. Antimicrobial susceptibility testing was done by Kirby-Bauer disk diffusion method and the results were interpreted according to the CLSI guidelines.Data obtained was analysed and presented in counts and percentages. 95 % confidence interval and p values were also calculated as applicable. Results: Imipenem was the most active antibiotic with 95.3 % susceptibility rate followed by Piperacillin-Tazobactem [89.7% ] and Levofloxacin [ 84.1% ] . Amikacin showed better susceptibility rate [74.7%] when compared to that of Gentamicin [65.4%]. The susceptibility rates to Cephalosporins and Aztreonam were relatively low. Most of the P.aeruginosa strains were isolated from clinical samples like pus [32.7% ],urine [ 26.1% ], and respiratory secretions [ 22.4% ]. 29 [27.1%] out of 107 clinical isolates of P.aeruginosa were found to be multidrug resistant. Most of the MDR P.aeruginosa strains were isolated from pus, urine, and respiratory secretions. Conclusions: The overall resistance demonstrated by clinical isolates of P.aeruginosa is high. This can be attributed to the inappropriate use of antibiotics. Guidelines on appropriate use of antibiotics not only reduces the morbidity and mortality in the patients infected with P.aeruginosa but also controls the emergence and spread of resistance among these pathogens.
Keywords: Antimicrobial agents, multidrug resistance, P. aeruginosa , susceptibility
Full Text:
INTRODUCTION
Pseudomonas aeruginosa is a very successful opportunistic pathogen. It can survive harsh environmental conditions and displays intrinsic resistance to a wide variety of antimicrobial agents that facilitates the organism’s ability to survive in the hospital setting. It is a notable cause of nosocomial infections of the respiratory and urinary tracts, wounds, blood stream, and even the central nervous system. In immunocompromised patients the infections are often severe and frequently life-threatening. In addition to its intrinsic resistance to various antibiotics, it also readily acquires resistance to the potentially active agents (1). Since some of the resistance markers are carried by promiscuous plasmids, the threat to human health is compounded by the possibility of transmission of the markers to other gram-negative pathogens (2).
Resistance to antipseudomonal antibiotics is increasing worldwide. This situation has been compounded by the lack of new classes of antipseudomonal drugs (3). Much of the antimicrobial resistance problem stems from the misuse of antibiotics, particularly excessive use. One of the main antibiotic resistance containement strategies is therefore to increase appropriate use and to reduce misuse, of antibiotics. Infection prevention and control activities to limit the spread of resistant bacteria are crucial (4). Area-wise studies on antimicrobial susceptibility profiles are essential to guide policy on the appropriate use of antibiotics. The present study was conducted to find out the antimicrobial susceptibility pattern of P.aeruginosa isolates obtained from various clinical samples at our hospital. The information would be useful in establishing empiric therapy guidelines and to contribute data to larger more extensive surveillance programs.
MATERIALS AND METHODS
The present study was conducted in the department of Microbiology, GSL Medical College and General Hospital, Rajahmundry, Andhra Pradesh, India. 107 isolates of P. aeruginosa were obtained from various samples received in the laboratory during the period January 2011 – June 2012. Ethical clearance was obtained from the institute. Routine standard operative procedures are followed in the laboratory in isolating and identifying the organisms from the clinical samples. P. aeruginosa was identified by Gram stain morphology, typical colony appearance, characteristic sweet grape like odour, blue-green appearance [pyocyanin pigment], motility and positive oxidase reaction (5). Antimicrobial susceptibility testing was done by Kirby-Bauer disk diffusion method and results are interpreted according to the Clinical and Laboratory Standards Institute (CLSI) guidelines (6). Antipseudomonas antibiotics like piperacillin (100 mcg), piperacillin-tazobactum (100/10 mcg), ceftazidime (30 mcg) , cefepime (30 mcg) , aztreonam (30 mcg), imipenem (10 mcg) , gentamicin (10 mcg) , amikacin (30 mcg), ciprofloxacin (5 mcg) and levofloxacin (5 mcg) were tested (HIMEDIA, MUMBAI, INDIA) . Multi-drug resistant (MDR) Pseudomonas aeruginosa strains were defined as isolates showing resistance to antibiotics from at least 3 of the 6 antipseudomonal classes of antimicrobial drugs tested in this study: antipseudomonal penicillins, cephalosporins, monobactams, carbapenems, aminoglycosides and fluoroquinolones. The data obtained in this study was summarized by counts and percentages. Antibiotic Susceptibility rates were presented with the respective 95% confidence interval values. Unpaired proportions were compared using Chi Square test or Fisher’s Exact Probability test, as appropriate . All analysis was two tailed and p < 0.05 was considered statistically significant.
RESULTS
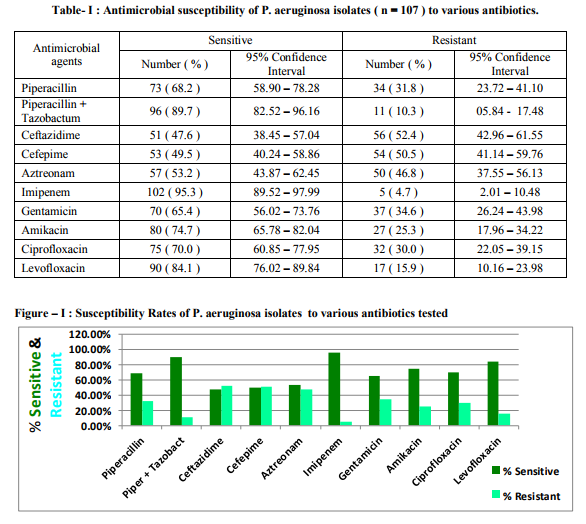
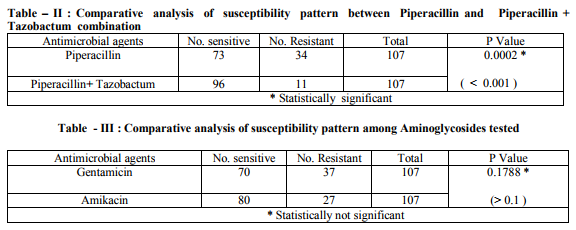
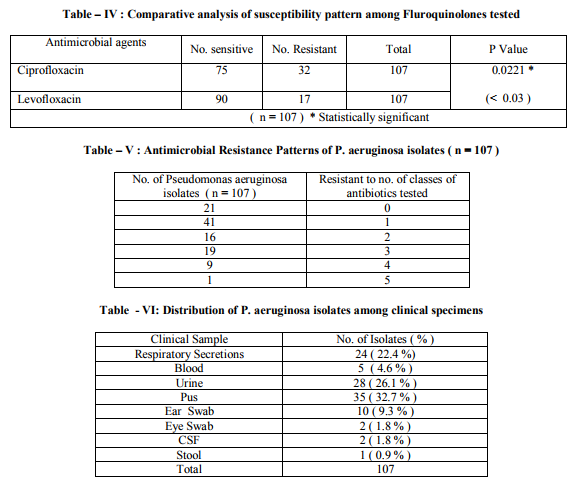
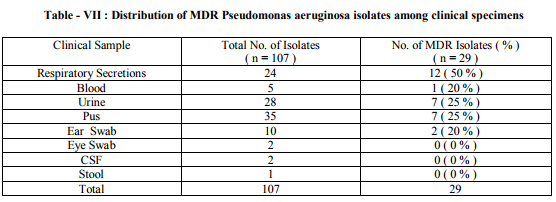
The results of antimicrobial susceptibility of P.aeruginosa clinical isolates to various antibiotics tested in this study are shown in Table – I and Figure - I . 95 % confidence interval data is also presented. Imipenem was the most active antibiotic with 95.3 % susceptibility rate. The next best were Piperacillin-tazobactam combination and levofloxacin with susceptibility rates of 89.7% and 84.1% respectively. The susceptibility rates between antibiotics that belonged to the same class were statistically compared as mentioned in Table – II , III and IV. Antimicrobial resistance patterns of P. aeruginosa isolates to various antibiotics are shown in Table – V. Out of 107 clinical isolates, 29 (27.1%) of P. aeruginosa were found to be multidrug resistant. The distribution pattern of P. aeruginosa isolates among various clinical samples is shown in Table -VI. Most of the P. aeruginosa strains were isolated from pus, urine, and respiratory secretions. The distribution pattern of MDR P.aeruginosa isolates among various clinical samples studied is shown in Table - VII. Most of the MDR P.aeruginosa strains were isolated from pus, urine, and respiratory secretions.
DISCUSSION
Antimicrobial surveillance should be done periodically to monitor the current susceptibility patterns in local hospitals (7) . In the present study, Imipenem was the most active antibiotic with 95.3 % susceptibility. This could be due to its restricted use in our hospital. This observation is in line with recent studies which reported very good sensitivity to carbapenems (8,9) However, some studies reported a notable resistance among the isolates of P.aeruginosa against carbapenems (7,10) . The second most active antibiotic was Piperacillin -Tazobactem with 89.7 % susceptibility. It had significantly better antibacterial activity against P.aeruginosa isolates than Piperacillin [68.2%] alone . This finding is in agreement with the study from Gujarat which has recommended the use of PiperacillinTazobactem like combinations against P. aeruginosa infections (10). Levofloxacin had better susceptibility rate [ 84.1% ] when compared to Ciprofloxacin [ 70.0% ]. This could be due to the overuse of Ciprofloxacin in our setting. The susceptibility rates to fluoroquinolones are better than previous Indian studies (11, 12). However they are in favour to the findings made in foreign studies (7, 13, 14). Among aminoglycosides Amikacin showed better susceptibility rate [74.7%] when compared to that of Gentamicin [65.4%] and it was found significant statistically. Better susceptibility rates to Amikacin were also reported by previous Indian studies (8,10 ) . They are generally used in combination with an antipseudomonal betalactam antibiotic. Amikacin use may be restricted to severe nosocomial infections (10). The rate of Susceptibility to Ceftazidime was 47.6 %. It was relatively low compared to the rates reported in various other studies (8,12,13,14) . This may be related to its frequent use in hospitalised patients where the possibility of emergence of resistance is high. However, the rate was better than that reported from Salem (11). Interestingly, the susceptibility rate to Aztreonam [53.2%] was not high even though it was not a commonly used antibiotic. Most of the P.aeruginosa strains were isolated from clinical samples like pus [ 32.7% ], urine [ 26.1% ], and respiratory secretions [ 22.4% ] . Out of 107 clinical isolates of P. aeruginosa , 29 [ 27.1% ] were multidrug resistant [ MDR ]. Most of these were isolated from pus, urine, and respiratory secretions. This indicates that infections with most resistant strains of P.aeruginosa are frequently encounterd in ICU units and post operative wards. This could be due to the frequent use of multiple antibiotics in critical care units. High prevalence rates of MDR P.aeruginosa strains in respiratory secretions was also reported by studies from Salem and Malaysia (7,12) . However, a standard definition of P.aeruginosa multidrug resistance would allow better comparisons between studies (7).
CONCLUSION
The overall resistance demonstrated by clinical isolates of P.aeruginosa is high. This can be attributed to the inappropriate use of antibiotics. Our study should guide clinicians on appropriate use of antibiotics. This not only reduces the morbidity and mortality in the patients infected with P.aeruginosa but also controls the emergence and spread of resistance among these pathogens. Regular monitoring of the use of antibiotics helps in preserving the effectiveness of antibiotics.
ACKNOWLEDGEMENT
Authors acknowledge the immense help received from the scholars whose articles are cited and included in references of this manuscript. The authors are also grateful to authors / editors /publishers of all those articles, journals and books from where the literature for this article has been reviewed and discussed.
References:
1. Betty A Forbes, Daniel F. Sahm, Alice S. Weissfeld. Baily and Scott’s Diagnostic Microbiology. 11th edition .Mosby Inc. 2002 ; 389 , 394.
2. Albert Balows, Brian I. Duerden. Topley and Wilson’s .Vol-2; Systemic Bacteriology; 9th edition . Arnold .1998 ; 1099
3. Fauci, Braunwald, Kasper, Hauser, Longo, Jameson et al. Harrison’s Principles of Internal Medicine; 17th edition. 2008 ; vol 1 ; 954
4. WHO Library Cataloguing-in-Publication Data ; The evolving threat of antimicrobial resistance: Options for action ; 2012
5. Collee JG, Fraser AG, Barry P Marmion, Simmons A. Mackie and McCartney Practical Medical Microbiology; 14th Edition. Churchill Livingstone, London. 1996 ; 417-418
6. Clinical and Laboratory Standard Institute, 2012. Performance standards for antimicrobial susceptibility testing. Clinical and Laboratory Standards Institute, Wayne. 22nd Informational Supplement, 32(3).
7. Siva Gowri Pathmanathan, Nor Azura Samat, Ramelah Mohamed. Antimicrobial susceptibility of clinical isolates of Pseudomonas aeruginosa from a Malaysian Hospital. Malaysian Journal of Medical Sciences. 2009 ; 16 (2): 27-32
8. Shenoy S, Baliga S, Saldanha DR, Prashanth HV. Antibiotic sensitivity patterns of Pseudomonas aeruginosa strains isolated from various clinical specimens. Indian J Med Sci 2002;56:427-30
9. Raja NS, Singh NN. Antimicrobial susceptibility pattern of clinical isolates of Pseudomonas aeruginosa in a tertiary care hospital. J Microbiol Immunol Infect. 2007;40:45-49.
10. Javiya VA, Ghatak SB, Patel KR, Patel JA. Antibiotic susceptibility patterns of Pseudomonas aeruginosa at a tertiary care hospital in Gujarat, India. Indian J Pharmacol 2008;40:230-4
11. K.M. Mohanasoundaram. Antimicrobial resistance in pseudomonas Aeruginosa ; Journal of Clinical and Diagnostic Research. 2011 June, Vol-5(3): 491-494
12. Anupurba S, Bhattacharjee A, Garg A, Sen MR. Antimicrobial susceptibility of Pseudomonas aeruginosa isolated from wound infections. Indian J Dermatol 2006;51:286-8
13. Farida Anjum, Asif Mir. Susceptibility pattern of pseudomonas aeruginosa against various antibiotics. Afr. J. Microbiol. Res.2010 ; Vol. 4 (10), 1005-1012
14. E.O.K. Nwankwo , S.A. Shuaibu . Journal of Medicine and Biomedical Sciences, 2010 . ISSN: 2078-0273
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License