IJCRR - 8(23), December, 2016
Pages: 29-38
Print Article
Download XML Download PDF
RAPID DETECTION OF MULTI-DRUG RESISTANT TUBERCULOSIS USING DIRECT DRUG SUSCEPTIBILITY TESTING AND LINE PROBE ASSAY
Author: Pranali Medhekar, Nilma Hirani, Sarala Menon, Ameeta Joshi, Abhay Chowdhary
Category: Healthcare
Abstract:Introduction: Tuberculosis, particularly multi-drug resistant tuberculosis (MDR TB), is a leading cause of morbidity and mortality in developing countries. Conventional methods of DST have a long turnaround time of around 2-3 months. Rapid detection of MDR TB can be achieved by direct tests, liquid based indirect susceptibility tests and molecular line probe assays. This study aims to evaluate the utility of Direct DST and line probe assay (LPA) for rapid detection of MDR TB.
Methodology: A total of 510 sputum concentrates (smear 1+/ more) were subjected to Direct DST and LPA. Conventional DST was put up from the primary cultures of all 510 strains using economic variant of 1% proportion method, considered the gold standard.
Results: Direct DST showed 100% specificity and sensitivity of 98.7%, 97.3% and 97.5% respectively for detection of Isoniazid (INH), Rifampicin (RIF) resistance and for MDR strains. LPA showed specificity of 98.4%, 96.8%, 97% and sensitivity of 97.5%, 98.9% and 98.9% for detection of INH, RIF resistance and MDR strains respectively. The commonest mutation pattern detected in our region was S531L (MUT3) for rpoB gene locus signifying RIF resistance + S315T1 (MUT 1) for katG gene locus signifying high level INH resistance.
Conclusion: Direct DST can be used as a good alternative for rapid detection of MDR-TB in resource poor settings. LPA is definitely a good tool, but requires requisite laboratory infrastructure and trained man-power. From our study, rifampicin resistance can be considered as a surrogate marker of MDR TB.
Keywords: MDR TB, Direct DST, LPA
Full Text:
INTRODUCTION
Tuberculosis (TB) continues to be a leading cause of morbidity and mortality in developing countries [1]. Global efforts for TB control are being challenged by the steady increase in drug resistant TB, particularly multidrug resistant tuberculosis (MDR TB), defined as resistance to rifampicin (RIF) and isoniazid (INH) [2]. As per the WHO Global TB Report 2015, there were an estimated 480 000 (range: 360 000-600 000) new cases of MDR-TB globally in 2014 and MDR TB has been reported in 3.3% new and 20% previously treated cases, respectively. [3]
MDR-TB requires 18-24 months of treatment with expensive second line drugs, some of which are parenteral drugs. The cure rate is much lower at about 60% than for drug susceptible TB [4]. Therefore, it is crucial that MDR TB should be detected as early as possible and measures implemented to effectively control its spread.
Detection of MDR-TB could be achieved by phenotypic and genotypic methods of drug sensitivity testing. The phenotypic methods include: a) Conventional methods of solid drug susceptibility testing [5] for detection of MDR TB which include 1% proportion method, resistance ratio and absolute concentration method. These methods require growth from primary cultures of specimens on Löwenstein Jensen (LJ) medium. Hence, they have a long turnaround time of around 2-3 months. b) Direct tests for DST of mycobacteria in which decontaminated sputum specimens are directly inoculated on drug free and drug containing LJ media. This obviates the need to wait for growth on primary culture, thus, reducing the turnaround time by 4-6 weeks and can be applicable in a resource poor setting. Examples of direct tests are Direct DST [6] on drug containing media, Nitrate Reductase Assay [NRA] [7], Microscopic Observation Drug Susceptibility [MODS] [7, 8]. c) Liquid based indirect susceptibility tests such as MB/BACT, BACTEC and MGIT960 which have an improved turnaround time of about 25 -45 days, as compared to the solid conventional DST, but, these are costly and not available where the need is greatest.[7,8]
The genotypic methods for rapid detection of MDR-TB include molecular line probe assays (LPA) which have a turnaround time of 48-72hrs. But, require specific laboratory infrastructure, technical expertise and are expensive. [9]
This study aims to evaluate the utility of DST and LPA for early detection of MDR-TB and also to evaluate rifampicin resistance as a surrogate marker for MDR-TB.
.MATERIAL AND METHODS
This was a prospective study carried out in a tertiary care hospital. The study protocol was approved by institutional ethics committee.
Inclusion criteria: Clinically diagnosed pulmonary tuberculosis (PTB) cases with sputum smear 1+ and more
Exclusion criteria: 1. Sputum smear positive PTB cases less than 1+.
2. Smear negative PTB cases.
3. Extrapulmonary (+/-) specimens.
METHODOLOGY
Each patient was asked to submit two sputum samples (one early morning and one spot sample) as per RNTCP Guidelines [11]. These were then subjected to direct smears by Ziehl Neelsen staining [12] and graded according to RNTCP guidelines [11]. The sputum samples were decontaminated and concentrated using the N-acetyl- L- Cysteine (NALC) – NaOH Concentration Method.[12]
Direct Drug Susceptibility Test (Direct DST) [6]
The sputum deposits obtained by NALC –NaOH method were inoculated with a 5 mm loop (27 SWG) onto a pair of plain LJ and drug containing LJ media (INH 0.2 μg /ml; R-40 μg/ml) along with a Para nitro benzoic acid (PNB) slant to confirm the growth of M.tuberculosis. The slopes were incubated at 37oC and examined weekly for 8 weeks and the growth was recorded weekly.
The definition of resistance was based on the amount of growth seen on the drug-free medium. Thus, when the growth on the drug-free medium was 2+ or more, growth of 1+ or more on the drug containing medium was defined as resistance to the drug. When growth on the drug-free medium is 1+ or less (i.e. < 100 colonies), any growth on the drug-containing medium was considered to be an indication of resistance to that particular drug. For this purpose, the higher growth observed on the paired slopes were considered for interpretation.
Conventional DST by 1% proportion method (economic variant) [5]
Inoculum preparation : With a 3 mm loop, 4-5 mg ( loopful) of primary culture was taken and inoculum was prepared in a sterile Bijou bottle containing 2 ml sterile distilled water and 6 sterile glass beads of diameter 3mm. Turbidity of the bacterial suspension was matched with McFarland standard no. 1 (10 7 to 108 cfu/ml ) against a black background. This was the neat bacterial suspension standardized at 1 mg/ml. Further’ 2 log dilutions were prepared. A loopful (using a 3 mm calibrated loop) of each dilution was inoculated on drug free and drug containing media [Isoniazid: 0.2μg/ml, Ethambutol: 2μg/ml, Streptomycin: 4μg/ml and Rifampicin: 40μg/ml.]
The slopes were incubated at 37°C. The growth was read at 28 days and again at 42 days.
(Growth recorded as - Confluent growth = 3+; More than 100 colonies = 2+; Record actual number of colonies = 1-100 cols.)
Interpretation
First reading was taken at 28th day after inoculation. The actual number of colonies (up to 100 colonies on the slope) on drug containing and drug-free slopes were counted.
Proportion of resistant bacilli existing in the strain was obtained by dividing the number of colonies obtained on drug containing slopes by that on drug free slope. Below a certain value – the critical proportion (1%)– the strain is classified as sensitive; above that value, it is classified as resistant. The proportions were reported as percentages. [5]
Internal Quality Control
Along with each batch of samples to be tested, a culture of H37Rv (a standard strain of M.Tuberculosis) or a known all sensitive strain was also incorporated and the test was validated.
Line Probe Assay (LPA) [13]:
Line probe assay is a DNA strip technology using a cellulose acetate membrane strip for the detection of resistance to RIF and/or INH. The kit used at our center for LPA was GenoType MTBDR Plus version 1.0 manufactured by Hain Lifescience, Germany which detects resistance to RIF and/or INH in sputum samples (smear 1+ / more) and culture isolates.. The steps involved are DNA Extraction from sputum concentrates, Multiplex Amplification, reverse hybridization and interpretation of results.
DNA Extraction:
A 500 µl of the smear positive (1+ or more) decontaminated sputum specimen was used for the DNA extraction which involved centrifugation of the specimen at 10,000 g for 15 minutes and suspension of the pellet in 100 µl of molecular grade water. It was then subjected to heat killing at 95oC for 20 minutes followed by sonication for 15 minutes and finally centrifugation at 10,000g for 5 minutes. 70-80 µl of the supernatant was taken and deep frozen at -20 0C of which 5 µl was used for PCR.
Multiplex amplification:
The master mix was prepared in a DNA-free room and the DNA extract was added to the master mix in a PCR hood. Both these procedures were done in separate rooms following the restricted access and unidirectional work flow protocol. A master mix negative control was also included with every batch of master mix prepared.
The amplification was done using a thermo cycler, with a set programme for amplification i.e. different for direct smear positive sputum specimens and culture isolates
Reverse hybridization:
It involved the chemical denaturation of the amplicons, hybridization of the biotin-labeled amplicons to the single stranded membrane-bound probes on the cellulose acetate membrane strip. It was followed by stringent washing, addition of streptavidine-alkaline phosphate conjugate to the wells and an enzymatic reaction mediated by alkaline phosphatase was observed in the form of a coloured band following addition of substrate in those regions where the amplicon and probe had hybridized. After the final washing, the strips were removed from the wells, allowed to air dry and placed onto the sheet provided with the kit and interpreted.
Each run incorporated one positive control and two negative controls (1 extraction negative control + 1 master mix negative control).
Interpretation of Results
- Valid results can be interpreted by the presence of CC (conjugate control) and AC (amplification control) bands for each sample run. The presence of TUB band indicates that the organism belongs to M.tuberculosis complex.
- A mutation in the relevant gene (and resistance to the relevant drug) is signified by either an absence of wild type band and/or the presence of a mutant band for each gene. The rpoB, katG and inhA gene loci each have a control band which needs to be present in order to interpret the results.
- The presence of mutations in the rpoB gene locus predicts RIF resistance, in katG predicts high level INH resistance while in the inhA gene locus predicts low level INH resistance.
- For interpretation of results, the intensity of all the bands (except CC band) should be equal to or greater than the intensity of the AC band.
- In order for a batch of results to be valid, the negative control strip has to have a CC and AC band present, but no other bands.
- Presence of bands other than CC and AC bands in the NC indicates contamination and the run need to be repeated.
- The product insert was referred to for interpretation of banding patterns.
Statistical analysis:
Sensitivity, specificity, positive predictive value, negative predictive value of Direct DST and LPA was calculated considering 1% proportion method the gold standard.
RESULTS
A total of 531 smear positive (1+ and more) sputum specimens were collected from the patients attending the outpatient as well as inpatient departments of tertiary care hospital. Out of these 531 specimens, 9 specimens showed no growth and12 were contaminated on Direct DST, thus could not be processed further by 1% Proportion method. Hence these 21 specimens were excluded from data analysis, since the results of Direct DST and 1% Proportion method (gold standard test for comparison) were not available for them. Primary cultures of the remaining 510 specimens were then subjected to economic variant of 1% proportion method. Thus total 510 specimens (n=510) all of which were subjected to all 3 tests were included for analysis.
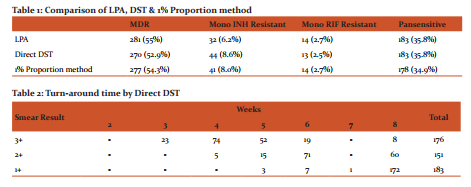
Table: 1 shows the comparative evaluation of LPA, Direct DST % 1% Proportion method in terms of the number of MDR, RIF and INH monoresistant and pansensitive strains.
Table: 2 shows the turn-around time by Direct DST. 74 out of 176 specimens with smear grading of 3+ gave results at 4th week, whereas for 172 out of 183 specimens with smear grading of 1+, results were obtained around 8th week; most of these being sensitive strains. Thus it was observed that higher the smear grading, we obtained the results earlier.
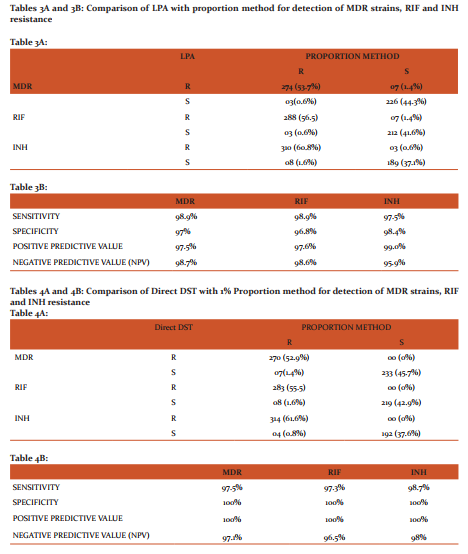
Tables: 3A and 3B- Comparison of LPA with proportion method for detection of MDR strains, RIF and INH resistance. LPA has shown a sensitivity of 98.9%, 98.9 % and 97.5 % and specificity of 97 %, 96.8 % and 98.4 % for detection of MDR, rifampicin and isoniazid resistance, respectively.
Tables: 4A and 4B- Comparison of Direct DST with 1% Proportion method for detection of MDR strains, RIF and INH resistance. DST has shown a sensitivity of 97.5%, 97.3 % and 98.7 % and specificity of 100% for detection of MDR, rifampicin and isoniazid resistance, respectively.
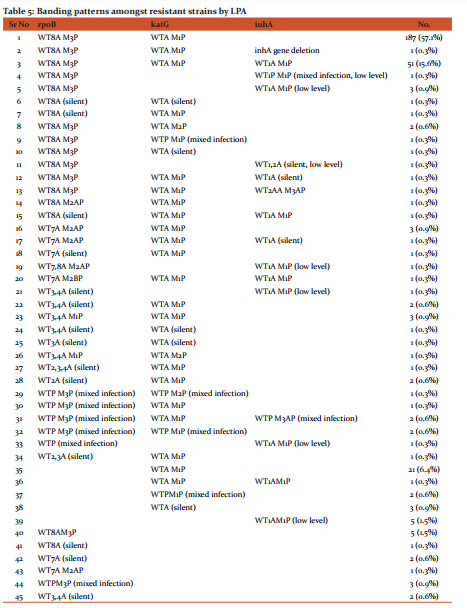
Table: 5 shows the banding patterns amongst resistant strains by LPA. There were 45 different patterns observed amongst the resistant strains. Commonest pattern (57.1%) seen was absence of wild type 8 band and simultaneous presence of mutation 3 band in the rpoB gene region coding for the S531L codon signifying RIF resistance + absence of wild type band and simultaneous presence of mutation 1 band in the katG gene region coding for S315T1 codon signifying high level INH resistance. 14 (2.7%) strains showed mixed infections and one strain showed complete inhA gene deletion. Low level INH resistance was seen in 13 (2.5%) of the 510 strains, of which 8 (1.5%) were MDR strains and 5 (0.9%) were INH mono-resistant strains.
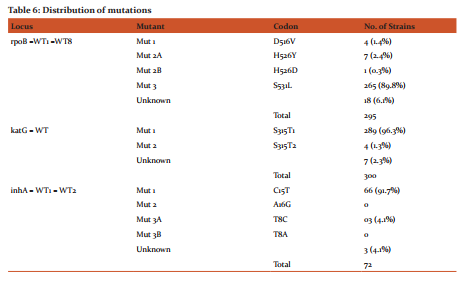
Table 6 shows the distribution of mutation in various gene loci. In the rpoB gene locus, the most common mutation was found to be the presence of MUT3 band coding for S531L codon in 265(89.8%) of the 295 strains. The katG gene locus showed maximum mutations in the MUT1 band region coding for S315T1 codon amounting to 289(96.3%) of the 300 strains, while in the inhA gene locus maximum mutations were seen in the MUT1 band region coding for C15T codon amounting to 66(91.7%) of the 72 strains.
DISCUSSION
In the present study, we have performed the comparative evaluation of the results of drug susceptibility testing of total 510 sputum specimens by Direct DST, Line probe assay (LPA), and 1% Proportion method, considered the gold standard.
The performance of the Direct DST and the Genotype MTBDR plus used for LPA test directly from smear positive (1+/ more) sputum specimens correlated very highly with conventional DST by 1% proportion method. (Table: 1)
The Direct DST described here would serve the purpose well in countries with limited resources. In our study, DST has shown a sensitivity of 97.5%, 97.3 % and 98.7 % and specificity of 100% for detection of MDR, rifampicin and isoniazid resistance, respectively. The concordance of the results of the 1% proportion method and Direct DST was found to be 97.6%. Thus, its performance characteristics suggest that the technique is equivalent to conventional 1% proportion method. The turn-around-time of the results obtained by Direct DST are shown in Table 2. Maximum results with smear grading of 3+ were obtained around 4th week, whereas for most of the specimens with smear grading of 1+ results were obtained around 8th week; most of these being sensitive strains. Thus it was observed that higher the smear grading, the results were obtained earlier.
The advantages of the Direct DST over the conventional 1% proportion method are that it gives sensitivity results at the same time as that of primary culture. This process not only reduces the turn-around-time by 4 weeks, but also contamination by eliminating the step of doing a subculture. Most importantly, the results of the Direct DST are more closely the representative of the bacterial population in the given sample unlike in the conventional test, which can suffer from errors of selection when drug susceptibility test is set up from a primary culture. In the Direct DST, resistance can be reported if adequate growth is seen on the drug slopes even when the plain medium is contaminated. Mathew et al also have reported the utility of Direct DST in MDR detection. Their results also indicate that resistance could be detected with growth as low as 10 colonies and in total agreement with the results of conventional test.[6] Hence, it is recommended that such results be accepted provisionally and confirmed with the conventional test on subculture.
The performance of Genotype MTBDRplus assay has co-related very well with the conventional 1% proportion method for DST with a sensitivity of 98.9%, 98.9 % and 97.5 % and specificity of 97 %, 96.8 % and 98.4 % for detection of MDR, rifampicin and isoniazid resistance, respectively. This test has the advantage of performing well and giving good results even in those specimens which are finally lost on culture.
Rifampicin resistance due to mutation in the rpoB gene was detected in 288 out of 291 phenotypically resistant RIF isolates. The resultant high sensitivity (98.9 %) for detection of rifampicin resistance found in our study could be attributed to all these mutations being present in the 81 bp- hotspot region of the rpoB gene (Table 6). This is in accordance with various other studies conducted across the world, such as S. Africa, Pakistan, Mongolia, India and Vietnam which have proven the mutations in the 81-bp region alone have got 98-100% sensitivity in detecting RIF resistance.[14-18]
Among all the gene mutations conferring to Rifampicin resistance, codon S531L was found to be the most frequently encountered mutation in 265 out of 295 (89.8%) which is comparable to the Indian study by Raveendran et al in 2012 (84.6%) [17], but, higher than that reported in various other studies across the world [14,16,18].
Mutations at codon 526 in our study were less common (2.7%) than reported from the studies conducted in various other studies such as S. Africa (8.6%)[14], Europe (15 %)[19], India (19 %)[17], Pakistan (22.5%)[15] and Iran (45. 6%)[20], but higher than the study conducted in Vietnam ( 1.8%)[18].
Mutations in the rpoB gene at the D516V codon were at 1.4 % in our study which is very low compared to the studies from S. Africa (9.6%)[14] and Europe (44%)[19].
18 isolates (6.1%) showed unknown mutations with the absence of both wild type and mutation band. It could be attributed to a rare mutation in the corresponding genomic region which is not probed on the strip (e.g mutations in the 530-533 region of the rpoB gene other than S531L mutation leading to absence of both, wild type 8 and MUT 3 which codes S531L codon specifically) or it could be a silent mutation since this test only screens the nucleic acid sequence and not the amino acid sequence. Therefore it is possible that mutations that do not cause an amino acid change (silent mutations) will still produce the absence of one of the wild type probes. In such cases phenotypic resistance determination should be considered[13]. If deletion of wild type band is the only indication of RIF resistance and there is no mutation probe, it would be advisable to continue the patient on RIF therapy till phenotypic sensitivity report is available.[17]
Mutations in the katG gene locus were detected in 300 out of 313 (95.8%) INH resistant strains. These were found to be maximum in the S315T1 codon region of the katG gene amounting to 96.3%. This is in accordance with Huyen et al’s study from Vietnam [18] and Van Rie et al’s study from S, Africa [21]. A high prevalence of katG mutations has been reported in high TB prevalent countries which is attributed to INH resistance and much lower prevalence in low TB-prevalent countries. This could be due to the on-going transmission of the strains in the high- burden settings.[22]
The presence of mutations in the inhA region signifying low level INH resistance accounts for 72 (23%) of the 313 INH resistant strains, with maximum mutations in the C15T (MUT1) region amounting to 66 (91.7%) of the 72 strains. Mutations in the inhA gene region accounting for 23.5% (66/281 strains) in MDRTB cases and 18.8% (6/32 strains) in INH mono-resistant cases were seen in our study, which is similar for MDR strains but lower for INH resistant strains when compared to the study by Barnard et al at 27% of MDR strains and 54% of INH mono-resistant strains.[14] The presence of mutations in inhA gene region vary in different geographical locations from 2% isolates in Western Cape Province of South Africa [21], 5.4% in the study by R. Raveendran et al[17] from India, 18% in the study from Vietnam by Huyen et al [18], 24% from KwaZulu-Natal [23] to 67.6% in Mongolia[16].
Out of the 327 strains showing mutations one strain exhibited complete inhA gene deletion, which is a very rare finding and the probable explanation for the absence of all inhA bands including the inhA locus control band, is a mutation in the primer annealing sequence, which results in missing of the specific amplification product. Since this strain was MDRTB with katG mutations as well, this presentation would not have any implication on patient treatment.
In our study, low level INH resistance alone was seen in 13 (2.5%) of the 510 strains, of which 8 (1.5%) were MDRTB strains and 5 (0.9%) were INH mono-resistant strains. These strains would not have been detected as MDR by the previous version of the assay (GenotypeMTBDR) which only tested for mutations in the katG gene.[24-26] The GenotypeMTBDRplus assay, with incorporation of additional probes to detect mutations in the inhA gene as well has a better sensitivity for detecting INH resistance.
In our study, 14 cases (2.7%) of mixed infections were detected by this assay showing presence of both the wild type as well as mutant band in the corresponding gene region. This is similar to the findings by Huyen et al[18] (3.6%). Of the 14 cases, mixed infections were seen in both, RIF and INH genomic loci in 5 (35.7%), while 9 (64.3%) out of 14 cases showed mixed infections either in the RIF or INH genomic loci.
In our study 21 discrepant results were obtained with LPA when compared to conventional DST, thus giving concordance of 95.9% (489/510 strains). 7/16 strains detected as RIF resistant by LPA were found to be sensitive on conventional DST. 5 out of these 7 false positive RIF resistant strains showed the presence of silent/unknown mutation which was not phenotypically expressed. In this situation results of phenotypic DST are confirmatory. For the rest of the false positive RIF resistant strains, performing phenotypic DST using minimum inhibitory concentration (MIC) would be confirmatory to check whether these isolates are showing resistance on a lower drug concentration. Rifampicin resistance was not detected in 3 strains by LPA which were RIF resistant on conventional DST. This could be attributed to other mutations outside the 81 bp-hotspot region which are not probed on the strip, hence LPA could not detect them. INH resistance was detected in 3 strains by LPA which were INH sensitive by conventional DST; 2 of which showed an unknown/silent mutation which was not phenotypically expressed and the remaining 1 strain showed low level INH resistance on LPA. LPA could not detect INH resistance in 8 strains which were resistant by conventional DST. This could be because of the mutation being present in other genomic loci such as kasA, intergenic region of oxyR-ahpC complex which are not probed on the DNA strip.[27,28]
This study also evaluated detection of monoresistance to INH and RIF. This approach was adopted because these 2 drugs are the key elements in short course TB chemotherapy and provide the most robust drug susceptibility results. Moreover, the only susceptibility data that are likely to be translated into clinical action (in terms of a change in drug regimen) in a programmatic setting would be identification of MDR TB. This is also seen with other well standardized methods, as resistance results for ETH and STR are less reliable than for INH and RIF. We found a higher incidence of INH monoresistance as compared to RIF monoresistance which is in accordance with the other studies worldwide. [14,16,18]
Because RIF mono resistance is rare, resistance to RIF is considered a surrogate marker of MDR TB. [29] Our results (Table 1) show that RIF resistance can be considered as a surrogate marker for MDR by all the three tests. Patients infected with RIF resistant strain of MTB generally have a poor prognosis, particularly because RIF resistance is often associated with resistance to other first line drugs.
Hence to summarize, successful treatment of MDR TB relies on prompt laboratory detection of drug resistance. This study found that Direct DST is a good rapid technique in resource-constraint settings. LPA is also a good rapid technique for rapid detection of MDR TB, but, it needs specific laboratory infrastructure, trained personnel, is expensive and needs an external quality assurance system. Both the techniques were well comparable with the gold standard 1% proportion method. Therefore, development of efficient laboratory strategies for rapid and reliable drug susceptibility testing of M tuberculosis is of importance for proper management of patients, particularly those with multi-drug resistant tuberculosis.
CONCLUSIONS
The results of this study have led us to following conclusions.
- Direct DST can be used as a good method, extremely feasible for rapid drug susceptibility testing of M tuberculosis in resource poor settings. Similarly, Direct DST can also compete with LPA in terms of turnaround time (as cheaper technique and less turnaround time compared to1% proportion method).
- Line probe assay is definitely a good tool for rapid detection of MDR provided one has the requisite laboratory infrastructure and technical expertise.
- 1% Proportion method still continues to remain the gold standard, when it comes to smear negative cases as well as to identify extensively drug resistant (XDR) cases of tuberculosis.
- Rifampicin resistance can be considered as a surrogate marker for MDR-TB in settings with high prevalence of drug resistant tuberculosis.
ACKNOWLEDGEMENT
We acknowledge the support from technical staff of TB culture and DST laboratory, Sir JJ Group of Hospitals Mumbai. We are also grateful to authors, editors and publishers of all those articles, journals and books included in the references of this manuscript.
References:
1. WHO: Global tuberculosis control 2008: surveillance, planning, financing, Geneva, Switzerland: World Health Organisation; Publication no, WHO/HTM/TB/2008.393, 2008.
2. XDR-TB (Extensively Drug Resistant Tuberculosis). WHO report 2006.
3. WHO: Global Tuberculosis Report, 2015
4. Blumberg HM, Burman WJ. Chaisson RE , Dailey CL, Etkind SC ,Fujivara P, et al: American Thoracic Society/ Centers For Disease Control and Prevention/ Infectious Diseases Society of America; treatment of Tuberculosis . Am. J Resp Crit Care Med 2003, 167(4) :603-662.
5. Manual of Standard Operating Procedures (SOPs). Culture of Mycobacterium tuberculosis and Drug Susceptibility Testing on solid medium. 1-142(2009) available at www.tbcindia.org
6. Mathew S, Nalini SM, Rahman F, Sundaram V and Paramasivan CN. Simple Direct Susceptibility Tests on sputum samples for early detection of resistance in tubercle bacilli: Indian J Tuber 2007; 54: 184-189.
7. Angeby KA , Klintz L, Hoffner SE :Rapid And Inexpensive Drug Susceptibility Testing of Mycobacterium tuberculosis with a nitrate reductase assay, J Clin Microbiol 2008 ,40(2) :553-555
8. Woods GL Susceptibility Testing for Mycobacteria Clin inf Dis 2000, 31 (5);209-1215.
9. Miotto P, PianaF, Cinllo DM, Miglori GB; Genotype MTBDRplus: a further step Toward rapid identification of drug resistant Mycobacterium tuberculosis J Clin Microbiol 2008, 46(1); 393-394.
10. Sharma M, Sethi S, Mishra B, Sengupta C, Sharma SK; Rapid detection of mutations in rpoB gene of rifampicin resistant Mycobacterium tuberculosis strains by line probe assay;Indian J Med Res 2003;pp 76-80.
11. Revised National Tuberculosis Control Programme: Guidelines on Programmatic Management of Drug Resistant TB (PMDT) in India May 2012.
12. Koneman’s Colour Atlas and Textbook of Diagnostic Microbiology. Washington Winn, Jr. Stephen Allen, William Janda, Elmer Koneman Gary Procop, Paul Schreckenberger, Gail Woods. 2006; Sixth Edition.
13. GenoType MTBDRplus,version 1.0 (product insert). (Internet) Feb 2007 (updated 2007 Apr 3; accessed 2007 May 9). Nehren, Germany; Hain Lifescience, GmbH. Available from: http:// www.hain-lifescience.com/pdf/304xx_pbl.pdf
14. Bernard N, Albert H, Coetzee G Rapid Molecular screening of multi drug resistant Tuberculosis in A high volume public health laboratory in South Africa; Am J Resp Cri Care Med, 2008,pp 787-792. 35 Int J Cur Res Rev | Vol 8 • Issue 23 • December 2016 Medhekar et.al.: Rapid detection of multi-drug resistant tuberculosis using direct drug susceptibility testing and line probe assay
15. Farooqui J. Q. Erun Khan, Syed Muhammed Zaheer Alam, Asho Ali, Zahra Hasan, Rumina Hasan, Line Probe Assay for detection of rifampicin and isoniazid resistant tuberculosis in Pakistan; Journal of Pakistan Medical Association; vol 62 No. 8, August 2012.
16. Buyankhishig B, T. Oyuntuya, B. Tserelmaa, J. Sarantuya, Marilla G. Lucero, S. Mitarai. Rapid molecular testing for multiresistant tuberculosis in Mongolia: A diagnostic study; International Journal of Mycobacteriology , vol 1, Issue 1, March 2012i
17. Raveendran R, Wattal C, Oberoi JK, Goel N, Datta S, Prasad KJ. Utility of Geno Type MTBDRplus assay in rapid diagnosis of multidrug resistant tuberculosis at a tertiary care centre in India; Indian Journal of Medical Microbiology vol 30, Issue 1, 2012.
18. Mai NT Huyen, Edine W Tiemersma, Nguyen TN Lan, Frank GJ Cobelens, Nguyen H Dung, Dinh N Sy, et al: Validation of the GenoType® MTBDRplus assay for diagnosis of multidrug resistant tuberculosis in South Vietnam; BMC Infectious Diseases 2010, 10:149.
19. Miotto P, Saleri N, Dembele M, Ouedraogo M, Baooum G, Pinsi G, et al. Molecular detection of rifampicin and isoniazid resistance to guide chronic TB patient management in Burkina Faso. BMC Infect Dis 2009;9:142.
20. Bahrmand AR, Titov LP, Tasbiti AH, Yari S, Graviss EA,. Highlevel rifampin resistance correlates with multiple mutations in the rpo B gene of pulmonary tuberculosis isolates from Afganistan border of Iran. J Clin Microbiol 2009; 47: 2744-50.
21. Van Rie A, Warren R, Mshanga I,Jordaan A, van der Spuy GD, Richardson M, Simpson J, Gie RP, Enarson DA, Beyer’s N et al. Analysis for a limited number of gene codons can predict drug resistance of Mycobacteriun tuberculosis in a high incidence community. J Clin Microbiol 2001;39-636-641.
22. Mokrousov I, Narvaskaya O, Otten T, Limenschenko E, Steklova L, Vyshnevskiy B. High prevalence of katG Ser315Thr substitution among isoniazid resistant Mycobacterium Tuberculosis clinical isolates from Northwestern Russia, 1996-2001. Antimicrob Agents Chemother 2002;46:1417-1424.
23. Kiepiela P, Bishop KS, Smith AN, Roux L, York DF. Genomic mutations in the katG, inhA and ahpC are useful for the prediction of isoniazid resistance in M. Tuberculosis isolates from KWA Zulu Natal, South Africa. Tuber Lung Dis 2000;80:87-56
24. Miotto P, Piana F, Penati V, Canducci F, Migliori GB, Cirillo DM. Use of Genotype MTBDR assay for molecular detection of rifampicin and isonoazid resistance in Mycobacterium tuberculosis in clinical strains isolated in Italy. J Clin Micobiol 2006;44:2485-2491.
25. Hilleman D, Rusch-Gerdes S, Richter E. Application of Genotype MTBDR assay directly on sputum samples. Int J Tuberc Lung Dis 2006;10:1057-1059.
26. Somoskovi A, Dormandy J, Mitsani D, Rivenberg J, Salfinger M. Use of smear-positive samples to assess the PCR-based genotype MTBDR assay for rapid, direct detection of the Mycobacterium tuberculosis complex as well as its resistance to isoniazid and rifampicin. J Clin Microbiol 2006;44:4459-4463.
27. Somoskovi A, Parsons LM, Salfinger M, The molecular basis of resistance to isoniazid, rifampin and pyrizinamide in Mycobacterium tuberculosis. Resspir Res 2001; 2:164-8.
28. Ramaswamy S, Musser JM. Molecular genetic basis of antimicrobial agent resistance in Mycobacterium tuberculosis: 1998 update. Tuber Lung Dis 1998; 79:3-29.
29. Molecular Line Probe Assays for Rapid Screening of Patients at Risk of Multi-Drug Resistant Tuberculosis (MDR-TB): WHO Expert group report.
|






 This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License